Research Article
Therapeutic Potential of Sand Box and In-Vitro Biochemical and Antimicrobial Potentials, Metallic Nanoparticle, and Molecular Docking-Based Screening of Key Phytochemicals
- F. A. Adewumi 1*
- M. O. Oseni 2
- V. O. Madu 3
- A. I. Airaodion 4
- G-M.M. Akinleye 5
- O. A. Ekundayo 1
- O. A. Oseni 6
1Department of Medical Laboratory Science, Faculty of Basic Medical Sciences, College of Medicine, Ekiti State University, Ado-Ekiti, Nigeria.
2Department of Chemistry, Faculty of Science, Federal University Oye-Ekiti, Ekiti State, Nigeria.
3Department of Science Laboratory Technology, Faculty of Science, Ekiti State University, Ado-Ekiti, Nigeria.
4Department of Biochemistry, Faculty of Natural and Applied Sciences, Lead City University, Ibadan, Oyo State, Nigeria.
5Department of Medical Microbiology and Parasitology, College of Medicine, Ekiti State University, Ado-Ekiti, Nigeria.
6Department of Medical Biochemistry, Faculty of Basic Medical Sciences, College of Medicine, Ekiti State University, Ado-Ekiti, Nigeria.
*Corresponding Author: F. A. Adewumi, Department of Medical Laboratory Science, Faculty of Basic Medical Sciences, College of Medicine, Ekiti State University, Ado-Ekiti, Nigeria.
Citation: F.A. Adewumi, M.O. Oseni, V.O. Madu, A.I. Airaodion, G-M.M. Akinleye, et al. (2026). Therapeutic Potential of Sand Box and In-Vitro Biochemical and Antimicrobial Potentials, Metallic Nanoparticle, and Molecular Docking-Based Screening of Key Phytochemicals, International Journal of Biomedical and Clinical Research, BioRes Scientia Publishers. 6(3):1-14. DOI: 10.59657/2997-6103.brs.26.120
Copyright: © 2026 F. A. Adewumi, this is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Received: January 16, 2026 | Accepted: February 03, 2026 | Published: February 09, 2026
Abstract
Background: Hura crepitans (Sandbox tree) is a tropical plant widely used in traditional medicine despite reports of inherent toxicity. Scientific evidence supporting its nutritional value and therapeutic potential remains limited.
Objectives: This study evaluated the nutritional composition, antinutritional factors, mineral profile, phytochemical constituents, antioxidant capacity, antimicrobial activity, nanoparticle-enhanced bioactivity, and molecular docking potential of H. crepitans leaf extract.
Methods: Fresh leaves of H. crepitans were collected from Ekiti State University, Ado-Ekiti, Nigeria. Proximate, mineral, and antinutritional analyses were performed using standard analytical procedures. Phytochemical constituents were qualitatively and quantitatively assessed. Antioxidant activity was evaluated using DPPH, hydrogen peroxide, nitric oxide scavenging, and ferric reducing antioxidant power (FRAP) assays. Antimicrobial activity of the crude extract and biosynthesized metallic nanoparticles was assessed against selected bacterial and fungal pathogens. Nanoparticles were characterized using UV-visible spectroscopy and Fourier-transform infrared (FTIR) analysis. Molecular docking was conducted to evaluate interactions between selected bioactive compounds and the dipeptidyl peptidase-4 (DPP-4) protein.
Results: Proximate analysis revealed high crude fiber content, moderate protein and lipid levels, and substantial ash content, indicating appreciable nutritional and mineral value. Antinutrient concentrations were within tolerable limits. Phytochemical screening confirmed the presence of phenolics, flavonoids, alkaloids, tannins, saponins, anthraquinones, and terpenoids. The extract exhibited significant In vitro antioxidant activity across all assays. While the crude extract showed modest antimicrobial effects, nanoparticle formulations markedly enhanced antibacterial and antifungal activities. Spectroscopic analyses confirmed successful nanoparticle synthesis mediated by plant phytochemicals. Molecular docking revealed strong binding affinities between key bioactive compounds and DPP-4, suggesting potential inhibitory activity relevant to metabolic regulation.
Conclusion: The findings provide scientific validation for the traditional use of H. crepitans and demonstrate its potential as a source of nutritionally valuable and pharmacologically active compounds with antioxidant and antimicrobial properties. Despite its known toxicity, the leaf extract exhibited promising therapeutic potential under controlled experimental conditions, warranting further in vivo studies and safety assessments.
Keywords: hura crepitans; phytochemicals; antioxidant activity; antimicrobial efficacy; medicinal plant; toxic plant; biochemical screening
Introduction
The sandbox tree (Hura crepitans) is a tropical species in the family euphorbias, native to the rainforests of Central and South America and parts of the Caribbean. It is widely recognized for its explosive seed pods and thorn-covered trunk, features that have earned it local names such as “dynamite tree” and “monkey no-climb” [1].
Across indigenous and traditional health systems, different parts of the plant, including the bark, seeds, and latex have long been used to manage a variety of ailments. Ethnobotanical reports describe its application as a purgative, vermifuge, and treatment for skin disorders and inflammatory conditions [2]. These long-standing practices suggest the presence of bioactive compounds with real therapeutic potential.
Phytochemical investigations of Hura crepitans have revealed several classes of secondary metabolites, notably diterpenoids, triterpenoids, saponins, and phenolic compounds. These constituents are associated with antimicrobial, anti-inflammatory, and cytotoxic activities [3]. At the same time, the plant is known to be highly toxic. Its latex contains potent irritants such as huratoxin and phorbol esters, which can trigger severe gastrointestinal and dermatological reactions if improperly handled or consumed [4]. This dual nature of medicinal promise and toxicity underscores the importance of controlled scientific evaluation.
Materials and Methods
Plant Identification Authentication
Fresh leaves of Hura crepitans were collected in March 2025 from the premises of Ekiti State University, Ado-Ekiti, Nigeria. The sample was authenticated and deposited in the herbarium of the Department of Plant Science, Faculty of Science, Ekiti State University. Botanical identification was carried out and confirmed by Mr. Omotayo, Chief Technologist in the department. Proper authentication ensured the integrity and reproducibility of all subsequent biochemical, antioxidant, toxicological, antimicrobial, and nanoparticle synthesis analyses.
Preparation and Extraction of Leaf Material
The leaves were carefully detached, washed thoroughly under running tap water to remove debris and surface contaminants, and then air-dried for several days. Once fully dried, the leaves were milled into a fine powder. Twenty grams of the powdered material were mixed with 100 mL of distilled water and agitated continuously overnight to allow efficient extraction of soluble phytochemicals.
Phytochemical Analysis (Fresh Samples)
Qualitative screening of the aqueous extract was carried out to detect major phytochemical groups including flavonoids, alkaloids, saponins, phenolic compounds, and steroidal nuclei, following standard procedures described in the literature [5-7].
DPPH Free Radical Scavenging Assay
The antioxidant capacity of the extract was evaluated using the DPPH radical scavenging method as modified by Gyamfi et al. [8]. Aliquots (1.0 mL) of extract at concentrations of 20, 40, and 80 mg/mL were each mixed with 1.0 mL of 0.1 mM methanolic DPPH solution. The mixtures were vortexed and incubated in the dark at room temperature for 30 minutes, after which absorbance was measured at 516 nm. A reduction in absorbance indicated radical scavenging activity.
Ferric Reducing Antioxidant Power (FRAP)
The reducing power of the extract was assessed using a modified ferric reducing antioxidant power method [9]. Various concentrations (10-50 µg/mL) of the extract were mixed with sodium phosphate buffer and potassium ferricyanide, incubated at 50°C for 20 minutes, and treated with trichloroacetic acid. After centrifugation, the supernatant was combined with distilled water and ferric chloride. Absorbance was read at 700 nm. The percentage reducing power was calculated as:
% FRAP= [(Abs control - Abs sample) /Abs control] x 100
Total Phenolic Content
Total phenolics were determined using the Folin-Ciocalteu method [10]. The reaction mixture was incubated at 45°C for 40 minutes and absorbance measured at 700 nm. Gallic acid served as the standard, and results were expressed as mg gallic acid equivalent (GAE) per gram of dry extract.
Total Flavonoid Content
Flavonoid content was quantified using a colorimetric assay [11]. Absorbance was measured at 510 nm, and results were expressed as mg GAE per gram of dry extract based on a gallic acid calibration curve.
Vitamin C Estimation
Ascorbic acid content was determined by oxidation with bromine water, derivatization with DNPH, and spectrophotometric measurement at 520 nm. Concentrations were calculated from a standard ascorbic acid calibration curve and expressed as mg/g of sample.
Green Synthesis and Antimicrobial Analyses
Green synthesis of metallic nanoparticles was performed using Hura crepitans leaf extract as a reducing and stabilizing agent. This eco-friendly approach avoids harsh chemicals and supports sustainable nanotechnology development.
Materials and Equipment
Key equipment included a blender, analytical balance, heating mantle, Buchner funnel, centrifuge, UV-Visible spectrophotometer, FTIR spectrometer, autoclave, incubator, and standard microbiological tools.
Synthesis of Silver, Copper, and Zinc Oxide Nanoparticles
Ten milliliters of the plant extract were added dropwise to 90 mL of 1 mM solutions of silver, copper, and zinc oxide salts, respectively. A visible color change signaled nanoparticle formation. The mixtures were heated for extended periods, centrifuged, washed with distilled water, and oven-dried prior to characterization.
Characterization of Nanoparticles
Formation of nanoparticles was monitored by UV-Visible spectroscopy in the 300-800 nm range. FTIR analysis was used to identify functional groups responsible for reduction and stabilization.
Antimicrobial Activity
Antimicrobial testing was conducted against Escherichia coli, Streptococcus mutans, Staphylococcus aureus, Salmonella typhi, and Klebsiella pneumoniae using a double dilution method [12].
Molecular Docking and In Silico Analysis
Molecular docking was performed to explore interactions between selected phytochemicals from Hura crepitans and relevant human protein targets. Eighteen ligands including flavonoids, phenolic acids, terpenoids, and fatty acid derivatives were retrieved from PubChem and prepared using Open Babel for docking studies [13,14].
ADMET and Drug-Likeness Prediction
Pharmacokinetic and toxicity profiles were evaluated using SwissADME and pkCSM platforms. Parameters assessed included absorption, distribution, metabolism, excretion, and toxicity, such as hERG inhibition, hepatotoxicity, and mutagenicity.
Interaction Analysis
To elucidate the functional context of the identified protein targets, protein–protein interaction (PPI) networks were constructed using the STRING database (https://string-db.org/), in accordance with the approach described by Badkas et al. [15]. Gene identifiers were used as input to explore interaction networks, signaling pathways, co-expressed proteins, and functional modules. In addition, target gene network analysis was performed using the eXpression2Kinases (X2K) platform (https://maayanlab.cloud/X2K/). This analysis enabled transcription factor enrichment, kinase-target expansion, and hierarchical PPI network construction. Collectively, these network-based analyses provided insights into the broader biological relevance and regulatory mechanisms underlying the predicted drug-target interactions.
Results
As shown in Table 1, Hura crepitans leaf exhibited low moisture content (5.77 ± 0.04%), indicating good shelf stability. The ash content was relatively high (14.09 ± 0.07%), indicating a significant mineral composition. Crude fiber was the dominant component (30.09 ± 0.01%), followed by crude fat (19.52 ± 0.06%), suggesting potential nutritional and functional relevance. Moderate protein (13.29 ± 0.04%) and carbohydrate (17.24 ± 0.04%) contents further support the nutritional value of the leaf.
Table 1: Proximate Composition of Hura crepitans leaf.
| Sample | Moisture Content | Ash | Crude Fat | Crude Fiber | Crude Protein | CHO |
| Sand Box | 5.77±0.04 | 14.09±0.07 | 19.52±0.06 | 30.09±0.01 | 13.29±0.04 | 17.24±0.04 |
Antinutrient Composition
Table 2 shows that Hura crepitans leaf contains moderate levels of antinutrients: oxalate (2.56 ± 0.05 mg/100 g), phytate (9.05 ± 0.02 mg/100 g), tannin (6.12 ± 0.05 mg/100 g), and trypsin inhibitor (52.95 ± 1.35%). These values are within tolerable limits, suggesting limited interference with nutrient bioavailability under normal consumption.
Table 2: Antinutrient Composition of Hura crepitans leaf.
| Sample | Oxalate mg/100g | Phytate mg/100g | Tannin mg/100g | Trypsin Inhibitor % |
| Sand Box | 2.56±0.05 | 9.05±0.02 | 6.12±0.05 | 52.95±1.35 |
Mineral Composition of Hura crepitans Leaf
The mineral analysis of Hura crepitans leaves (Table 3) revealed that potassium (K) was the most abundant microelement at 5.32 ± 0.02 mg/100 g, followed by calcium (Ca, 6.98 ± 0.01 mg/100 g), magnesium (Mg, 3.12 ± 0.03 mg/100 g), and sodium (Na, 2.95 ± 0.01 mg/100 g). Among trace elements, iron (Fe) was present at 3.16 ± 0.00 ppm, copper (Cu) at 0.72 ± 0.00 ppm, manganese (Mn) at 0.32 ± 0.00 ppm, and chromium (Cr) at 0.23 ± 0.00 ppm. Lead (Pb) was detected at a low concentration of 0.12 ± 0.00 ppm, indicating minimal toxic risk. These results demonstrate that the leaves of H. crepitans contain essential macro- and microelements that can contribute to nutritional and physiological functions.
Table 3: Mineral Composition of Hura crepitans leaf.
| K mg/100 | Na mg/100 | Ca mg/100 | Mg mg/100 | Cu Ppm | Fe Ppm | Mn Ppm | Cr Ppm | Pb Ppm |
| 5.32±0.02 | 2.95±0.01 | 6.98±0.01 | 3.12±0.03 | 0.72±.000 | 3.16±0.00 | 0,32±0.00 | 0.23±0.00 | 0.12±0.00 |
Phytochemical Screening of Aqueous Extract
Phytochemical analysis (Table 4) revealed a rich presence of bioactive compounds in the aqueous ext act of the plant. Phenolics, terpenoids, and cardiac glycosides were strongly detected (++++), while alkaloids (Wagner’s test) and anthraquinones were moderately present (+++). Saponins showed moderate occurrence (++), whereas steroids (Liebermann’s test) were detected in low abundance (+). Overall, the results indicate that the extract is rich in phytochemicals associated with antioxidant and therapeutic activities.
Table 4: Phytochemical screening of the Aqueous plant’s extracts.
| Saponin Test | ++ |
| Phenolic Test | ++++ |
| Wagner’s Test | +++ |
| Anthraquinone Test | +++ |
| Terpenoid Test | ++++ |
| Keller-killani Test | ++++ |
| Lieberman’s Test | + |
Antioxidant Potential of Aqueous Extract
The aqueous leaf extract of Hura crepitans exhibited appreciable antioxidant potential (Table 5). The extract contained a high flavonoid content (54.29 ± 0.02 mg AAE/g) and a substantial total phenolic content (37.00 ± 0.01 mg GAE/g), indicating a strong pool of polyphenolic antioxidants. Vitamin C was present at a lower level (1.97 ± 0.002 mg/g), suggesting that the antioxidant activity of the extract is largely attributable to flavonoids and phenolic compounds rather than ascorbic acid.
Table 5: Antioxidant potentials of aqueous plant’s extracts of Sand box Hura crepitans leaf.
| Sample | Flavonoid Content (mg AAE/g) | Phenolic Content (mg GAE/g) | Vitamin C Content |
| Sand Box | 54.29±0.02 | 37.00±0.01 | 1.97±0.002 |
Free Radical Scavenging Activity
The aqueous leaf extract of Hura crepitans demonstrated notable free radical scavenging activity across multiple In vitro assays (Table 6). The extract exhibited strong hydrogen peroxide scavenging activity (80.00 ± 0.03%), moderate ferric reducing antioxidant power (66.00 ± 0.01%), and appreciable DPPH (57.00 ± 0.03%) and nitric oxide (50.00 ± 0.02%) scavenging activities. These results indicate a broad-spectrum antioxidant capacity, reflecting the ability of the extract to neutralize different reactive species through both radical scavenging and reducing mechanisms.
Table 6: % Free radicals scavenging of Sand box Hura crepitans leaf.
| Sample | DPPH | H2O2 | NO | FRAP |
| Sand Box | 57.00±0.03 | 80.00±0.03 | 50.00±0.02 | 66.00±0.01 |
Antibacterial Activity
The antibacterial activity of the aqueous leaf extract of Hura crepitans and its biosynthesized nanoparticles against selected bacterial pathogens is presented in Table 7a. The crude aqueous extract exhibited low to modest antibacterial activity, with inhibition zones ranging from 2.00 to 4.00 mm across the tested organisms. The highest activity of the extract was observed against Salmonella typhi and Klebsiella pneumoniae (4.00 mm each), while the least inhibition was recorded against Streptococcus mutans (2.00 mm). In contrast, nanoparticle formulations markedly enhanced antibacterial efficacy. Silver nanoparticles (AgNPs) produced inhibition zones of 3.00-11.00 mm, with pronounced activity against Staphylococcus aureus (11.00 mm). Zinc nanoparticles (ZnNPs) showed improved and more uniform activity (5.00-8.50 mm), particularly against Escherichia coli and Klebsiella pneumoniae. Copper nanoparticles (CuNPs) exhibited the strongest antibacterial effect overall, with inhibition zones ranging from 8.00 to 11.00 mm, demonstrating high potency against both Gram-positive (Staphylococcus aureus, Streptococcus mutans) and Gram-negative (E. coli, Salmonella typhi, K. pneumoniae) bacteria. Overall, the results indicate that nanoparticle incorporation significantly enhances the antibacterial activity of H. crepitans leaf extract, with copper nanoparticles showing the greatest broad-spectrum antibacterial potential. These findings support the application of plant-mediated metallic nanoparticles as effective antimicrobial agents.
Table 7a: Antibacterial activity of the aqueous extract and the Nanoparticle of Sandbox Hura crepitans leaf on some selected bacteria.
| Sample | E. coli | Streptococcus mutant | Salmonella typhi | Staphylococcus aureus | Klebsiella pneumoniae |
| Zone Inhibition in mm | |||||
| Sand Box Extract | 3.50 | 2.00 | 4.00 | 3.00 | 4.00 |
| AgNP | 7.00 | 3.00 | 5.20 | 11.00 | 7.00 |
| ZnNP | 8.50 | 6.00 | 5.50 | 5.00 | 8.00 |
| CuNP | 8.60 | 8.00 | 8.00 | 11.00 | 9.00 |
Antifungal Activity
The antifungal activity of the aqueous leaf extract of Hura crepitans and its biosynthesized nanoparticles against Macrophomina phaseolina and Alternaria alternata is summarized in Table 7b. The crude aqueous extract exhibited moderate antifungal activity, with mycelial growth inhibition of 40.16% against M. phaseolina and 41.05% against A. alternata. Nanoparticle formulations demonstrated enhanced antifungal efficacy compared with the crude extract. Silver nanoparticles (AgNPs) increased mycelial growth inhibition to 44.21% and 46.84% against M. phaseolina and A. alternata, respectively. Zinc nanoparticles (ZnNPs) showed strong inhibitory activity, particularly against A. alternata (54.74%), representing the highest inhibition recorded for this fungus. Copper nanoparticles (CuNPs) exhibited the greatest activity against M. phaseolina (53.00%) and also showed substantial inhibition against A. alternata (52.63%). Overall, the results demonstrate that nanoparticle incorporation significantly improves the antifungal potential of H. crepitans leaf extract, with CuNPs and ZnNPs showing superior activity against the tested phytopathogenic fungi. These findings highlight the promise of plant-mediated metallic nanoparticles as effective antifungal agents for biomedical and agricultural applications.
Table 7b: Antifungal activity of Aqueous extract and the Nanoparticle of Sand box Hura crepitans leaf on some selected bacteria.
| Samples | Microphomina phaseolina | Alternaria alternata |
| Mycelia Growth Inhibition (%) | ||
| Sand box extract | 40.16 | 41.05 |
| AgNP | 44.21 | 46.84 |
| ZnNp | 46.84 | 54.74 |
| CuNP | 53.00 | 52.63 |
Ligand Identification and Preparation
Table 8 presents the major phytochemical constituents of Hura crepitans leaf identified and selected for ligand preparation in the molecular docking analysis. A total of eighteen compounds representing diverse chemical classes, including organic acids, aldehydes, phenolics, fatty acids, amides, heterocyclic compounds, and steroids, were retrieved from the PubChem database using their respective compound identification (CID) numbers. The canonical SMILES strings were used to define the molecular structures and ensure accurate three-dimensional representation during docking simulations. The identified ligands included low-molecular weight compounds such as acetaldehyde, acetic acid, propanoic acid, and dihydroxyacetone, as well as phenolic and aromatic compounds including 4H-pyran-4-one, 3-furaldehyde, 1,2-benzenediol, m-toluylic acid, and gallic acid. Higher molecular weight bioactive constituents such as long-chain fatty acids (n-hexadecanoic acid, octadecanoic acid, and 9,12-octadecadienoic acid), esters, amides, and terpenoid/steroidal compounds (squalene and estrone) were also included. The structural diversity of the selected ligands reflects the complex phytochemical profile of H. crepitans and provides a robust basis for evaluating multiple modes of interaction with the target protein. These prepared ligands were subsequently used for molecular docking studies to assess their binding affinities and potential biological relevance.
A total of eighteen phytochemical compounds were identified from Hura crepitans leaf extract and prepared as ligands for In Silico analyses (Table 8). The compounds comprised low–molecular weight organic acids and aldehydes (acetaldehyde, acetic acid, propanoic acid), phenolic and heterocyclic compounds (4H-pyran-4-one, 3-furaldehyde, 1,2-benzenediol, gallic acid), fatty acids and their derivatives (9,12-octadecadienoic acid, n-hexadecanoic acid, octadecanoic acid), amides, esters, terpenoids, and steroidal constituents such as estrone and squalene. Canonical SMILES and PubChem compound IDs were used to ensure structural accuracy prior to docking simulations.
Table 8: Ligand Preparation Identification of major phytochemical constituents of Hura creptains.
| S/N | Names of Ligand | Cid No. | Cannonial Smiles |
| 1 | Acetaldehyde | 177 | CC=O |
| 2 | Acetic Acid | 176 | CC(=O)O |
| 3 | Propanoic acid | 1032 | CCC(=O)O |
| 4 | 4H-Pyran-4-one | 7968 | C1=COC=CC1=O |
| 5 | 3-Furaldehyde | 10351 | C1=COC=C1C=O |
| 6 | 1,2 benezendiol | 951 | C1=CC(=C(C=C1C(CN)O)O)O |
| 7 | m-Toluylic acid | 7418 | CC1=CC(=CC=C1)C(=O)O |
| 8 | Galic acid | 46780424 | C1=CC=C(C=C1)COC(=O)C2=CC(=C3C(=C2)OC(O3)(C4=CC=CC=C4)C5=CC=CC=C5)O |
| 9 | Dihydroxyacetone | 70709 | C(C(=O)CO)O |
| 10 | Nonanamide | 670 | CCCCCCCCC(=O)N |
| 11 | Octanoic acid, 3-hydroxy-, methyl ester | 110974 | CCCCCC(CC(=O)OC)O |
| 12 | Acetamide, N-tetrahydrofurfuryl-2-methoxy- | 532079 | COCC(=O)NCC1CCCO1 |
| 13 | 3-Vinyl-1,2-dithiacyclohex-4-ene | 525328 | C=CC1C=CCSS1 |
| 14 | 9,12-Octadecadienoic acid (Z,Z)- | 5280450 | CCCCC/C=C\C/C=C\CCCCCCCC(=O)O |
| 15 | n-Hexadecanoic acid | 985 | CCCCCCCCCCCCCCCC(=O)O |
| 16 | Estrone | 5870 | C[C@]12CC[C@H]3[C@H]([C@@H]1CCC2=O)CCC4=C3C=CC(=C4)O |
| 17 | Octadecanoic acid | 5281 | CCCCCCCCCCCCCCCCCC(=O)O |
| 18 | Squalene | 638072 | CC(=CCC/C(=C/CC/C(=C/CC/C=C(/CC/C=C(/CCC=C(C)C)\C)\C)/C)/C)C |
In Silico Target Prediction and ADME Profiling
In Silico target prediction revealed that the identified phytochemicals interact with multiple biologically relevant protein targets (Table 9). Notably, 1,2-benzenediol showed high prediction probabilities for several G-protein-coupled receptors, including α- and β-adrenergic receptors and dopamine D2 receptors. Fatty acid-derived compounds such as 9,12-octadecadienoic acid and n-hexadecanoic acid demonstrated strong associations with fatty acid binding proteins (FABP2, FABP3, FABP4, FABP5) and nuclear receptors including peroxisome proliferator-activated receptors (PPARα, PPARδ, and PPARγ).
Table 9: The In Silico ADME (absorption, distribution, metabolism, and excretion) screening of the compounds.
| Compound Name | Target | Common name | Uniprot ID | ChEMBL ID | Target Class | Probability | Known actives (3D/2D) |
| 1,2 benezendol | Alpha-2a adrenergic receptor | ADRA2A | P08913 | CHEMBL 1867 | Family A G protein - couplled receptor | 0.999408 | 4/7 |
| Adrenergic receptor alpha -2 | ADRA2C | P18825 | CHEMBL 1916 | Family A G protein - couplled receptor | 0.8275 | 2/5 | |
| Alpha-2b Adrenergic receptor (by homology) | ADRA2B | P18089 | CHEMBL 1942 | Family A G protein - couplled receptor | 0.8275 | 3/7 | |
| Adrenergic receptor beta | ADRD2 | P07550 | CHEMBL 210 | Family A G protein - couplled receptor | 0.8275 | 7/69 | |
| Beta -1 adrenergic receptor (by homology) | ADRB1 | P08588 | CHEMBL 213 | Family A G protein - couplled receptor | 0.8275 | 12/32 | |
| Dopamine D2 receptor | DRD2 | P14416 | CHEMBL 214 | Family A G protein - couplled receptor | 0.8275 | 5/139 | |
| Alpha-1a adrenergic receptor | ADRA1A | P35368 | CHEMBL 229 | Family A G protein - couplled receptor | 0.8275 | 6/11 | |
| 9,12 Octadecadienoic | Fatty acid binding protein adipocyte | FABP4 | P15090 | CHEMBL 2083 | Fatty acid binding protein family | 1 | 26/4 |
| Peroxisome proliferator-activated receptor gamma | PPARG | P37231 | CHEMBL 235 | Nuclear receptor | 1 | 364/22 | |
| Peroxisome proliferator-activated receptor alpha | PPARA | Q07869 | CHEMBL 239 | Nuclear receptor | 1 | 225/18 | |
| Fatty acid binding protein in muscle | FABP3 | P05413 | CHEMBL 3344 | Fatty acid binding protein family | 1 | 15/5 | |
| Cyclooxygenase-1 | PTGS1 | P2321 | CHEMBL 221 | Oxidoreductase | 0.794118 | 4/2 | |
| Free fatty acid receptor 1 | FFARD1 | Q14842 | CHEMBL 4422 | Family A G protein - couplled receptor | 0.794118 | 168/3 | |
| Peroxisome proliferator-activated receptor delta | PPARD | Q03181 | CHEMBL 3979 | Nuclear receptor | 0.775875 | 177/10 | |
| n-hexadecanoic acid | Fatty acid binding protein adipocyte | FABP4 | P15090 | CHEMBL 2083 | Fatty acid binding protein family | 0.935895 | 20/30 |
| Peroxisome proliferator-activated receptor alpha | PPARA | Q07869 | CHEMBL 239 | Nuclear receptor | 0.935895 | 160/9 | |
| Fatty acid binding protein muscle | FABP3 | P05413 | CHEMBL 3344 | Fatty acid binding protein family | 0.935895 | 9/5 | |
| Fatty acid binding protein epidermal | FABP5 | Q01469 | CHEMBL 3674 | Fatty acid binding protein family | 0.935895 | 3/2 | |
| Peroxisome proliferator-activated receptor delta | PPARD | Q0318 | CHEMBL 3979 | Nuclear receptor | 0.935895 | 148/7 | |
| Fatty acid binding protein intestinal | FABP2 | P12104 | CHEMBL 4879 | Fatty acid binding protein family | 0.935895 | 1/1 | |
| Free fatty acid receptor 1 | FFAR1 | O14842 | CHEMBL 4422 | Family A G protein - couplled receptor | 0.598213 | 153/3 | |
| Estrone | Serotonin 2b(5-H2b) | HTR2B | P41595 | CHEMBL 1833 | Family A G protein - couplled receptor | 1 | 2/5 |
| Androgen receptor (by homology) | AR | P10275 | CHEMBL 1871 | Nuclear receptor | 1 | 43/34 | |
| Cytochrome P45019A1 | CYP19A1 | P11511 | CHEMBL 1978 | Cytochrome P450 | 1 | 150/75 | |
| Estrogen receptor alpha | ESR1 | P03372 | CHEMBL 206 | Nuclear receptor | 1 | 118/121 | |
| Serotonin transporter | SLC6A4 | P31645 | CHEMBL 228 | Electrochemical transporter | 1 | 4/45 | |
| Estrogen receptor beta | ESR2 | Q92731 | CHEMBL 242 | Nuclear receptor | 1 | 115/111 | |
| Estradiol 17 beta dehydrogenase 1 | HSD17B1 | P14061 | CHEMBL 3181 | Enzyme | 1 | 35/48 | |
| Testis-special androgen binding protein | SHBG | P04278 | CHEMBL 3305 | Secreted protein | 1 | 9/18 | |
| Octadecadienoic acid | Peroxisome proliferator-activated receptor | PPARA | Q07869 | CHEMBL 239 | Nuclear receptor | 0.929299 | 115/9 |
| Peroxisome proliferator-activated receptor delta | PPARD | Q03181 | CHEMBL 3979 | Nuclear receptor | 0.929299 | 135/7 | |
| Fatty acid binding protein adipocyte | FABP4 | P15090 | CHEMBL 2083 | Fatty acid binding protein family | 0.714850 | 13/3 | |
| Fatty acid binding protein epidermal | FABP5 | Q01469 | CHEMBL 3674 | Fatty acid binding protein family | 0.714850 | 3/2 | |
| Fatty acid binding protein muscle | FABP3 | P05413 | CHEMBL 3344 | Fatty acid binding protein family | 0.526361 | 9/5 | |
| Fatty acid binding protein intestinal | FABP2 | P12104 | CHEMBL 4879 | Fatty acid binding protein family | 0.526361 | 1/1 |
Estrone exhibited strong predicted interactions with multiple nuclear and endocrine-related receptors, including estrogen receptors (ESR1 and ESR2), androgen receptor, serotonin receptors, cytochrome P450 enzymes, and steroid metabolism proteins, all with high probability scores. These results indicate that H. crepitans phytochemicals may influence metabolic regulation, lipid homeostasis, inflammatory pathways, and hormone-related signaling mechanisms.
Molecular Docking Analysis
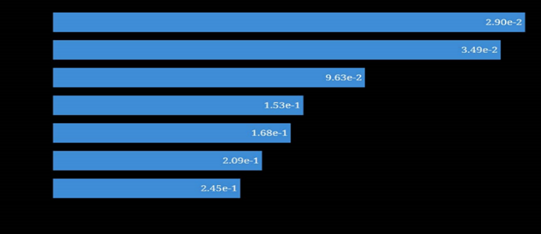
Molecular docking results demonstrated variable binding affinities between the prepared ligands and the target receptor (Table 10). Gallic acid showed the strongest binding affinity with a docking score of -8.9 kcal/mol, followed by estrone (-8.0 kcal/mol), indicating highly stable ligand-receptor interactions. Moderate binding affinities were observed for m-toluylic acid (-5.7 kcal/mol), 9,12-octadecadienoic acid (-5.2 kcal/mol), and n-hexadecanoic acid (-4.8 kcal/mol). Lower molecular weight compounds such as acetic acid, propanoic acid, and acetaldehyde exhibited weaker interactions. Overall, the docking results suggest that phenolic and steroidal constituents contribute most significantly to the biological activity of H. crepitans.
Table 10: Receptor binding affinities/free energy of binding in kcal/mol unit.
| Ligands | Docking scores/Affinity |
| Galic acid | -8.9 |
| Estrone | -8 |
| m-Toluylic acid | -5.7 |
| 9,12 Octadecadienoic acid | -5.2 |
| n-Hexadecanoic | -4.8 |
| Octanoic acid | -4.6 |
| Acetamide, N-tetrahydrofurfuryl-2-methoxy | -4.5 |
| 4H-Pyran-4-one | -4.4 |
| Nonanamide | -4.2 |
| 3-furaldehyde | -4.1 |
| 2, dihydroxyacetone | -4 |
| Dihydroxyacetone | -3.8 |
| Propanoic acid | -3.8 |
| Acetic acid | -3.4 |
| Acetaldehyde | -2.9 |
UV–Visible Spectroscopy and Antioxidant Quantification
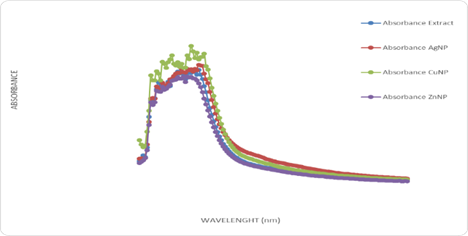
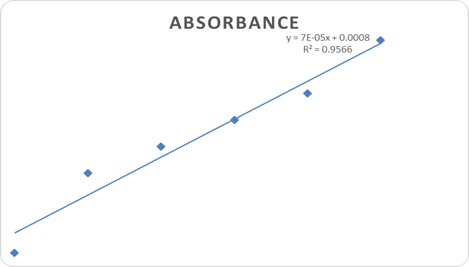
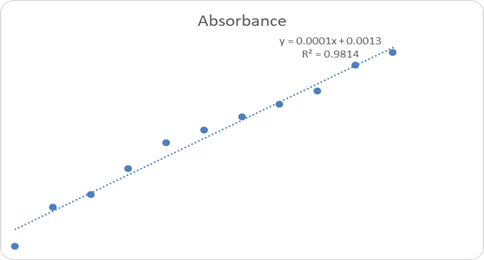
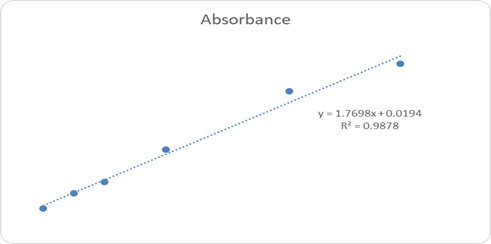
UV-visible spectroscopic analysis confirmed the successful interaction of H. crepitans aqueous extract with metallic nanoparticles, as shown by characteristic wavelength shifts and absorbance peaks (Figure 1). Quantitative antioxidant profiling revealed concentration-dependent increases in absorbance for flavonoids (Figure 2), phenolic compounds (Figure 3), and ascorbic acid (Figure 4), confirming the strong antioxidant potential of the extract.
Figure 1: The Wavelength scanning of Hura crepitans aqueous extract with metallic nanoparticles.
Figure 2: Flavonoid Concentration and Absorbance Graph.
Figure 3: Phenol Concentration and Absorbance Graph.
Figure 4: Ascorbic Acid Concentration and Absorbance Graph.
FTIR Characterization
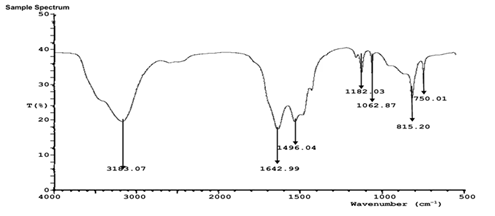
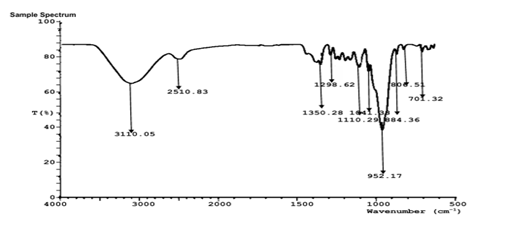
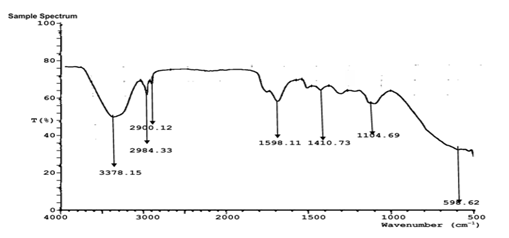
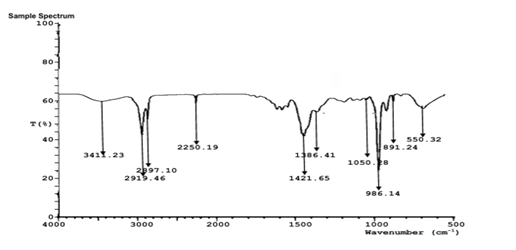
FTIR analysis of the aqueous extract and synthesized nanoparticles (Figures 5a-d) revealed distinct absorption bands corresponding to hydroxyl (-OH), carbonyl (C=O), amine (N-H), and alkyl (C-H) functional groups. These functional groups indicate the presence of polyphenols, proteins, and other biomolecules responsible for the reduction, stabilization, and capping of silver, zinc, and copper nanoparticles. The similarity of major functional peaks across extract and nanoparticle spectra confirms phytochemical-mediated nanoparticle synthesis.
Sample 1D: Sand box Hura crepitans plant aqueous extract
Resolution :8
Sample Scan: 200 scans
Full Scale 42243
Method: Transmittance Method
Figure 5a: FTIR characterization of Sand box (Hura crepitans) aqueous extract.
Figure 5b: FTIR characterization of Sand box (Hura crepitans) AgNP.
Figure 5C: FTIR characterization of Sand box (Hura crepitans) ZnNP.
Figure 5D: FTIR characterization of Sand box (Hura crepitans) CuNP.
Figure 6.1: Transcription Factor Enrichment Analysis (TFEA).
Figure 6.2: Kinase Enrichment Analysis (KEA).
Figure 6.3: eXpression2Kinases Network.
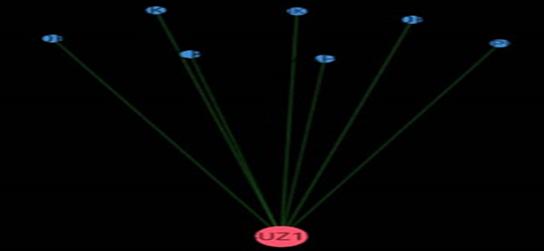
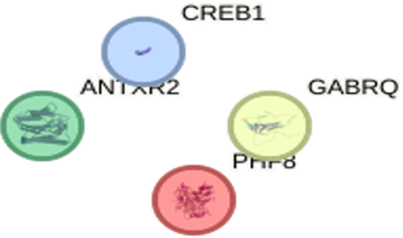
Figure 6.4: Protein- Protein interaction of Hura crepitans molecular Targets.
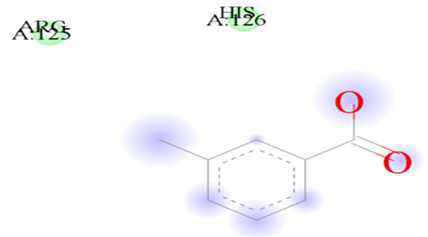
Figure 7.1: The binding of Galicacid against the target macro molecule (DPP4).
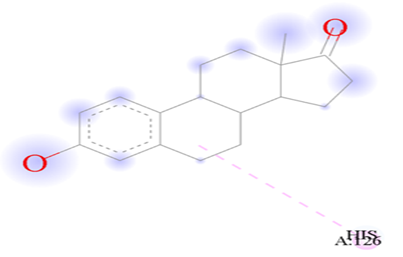
Figure 7.2: Showing the binding of Estrone against the target macro molecule (DPP4).
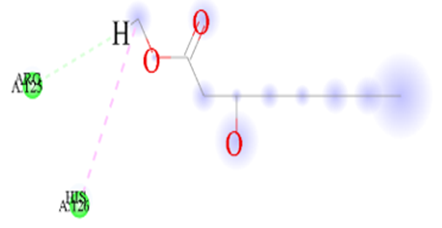
Figure 7.3: The binding of m-Toluylic_acid against the target macro molecule (DPP4).
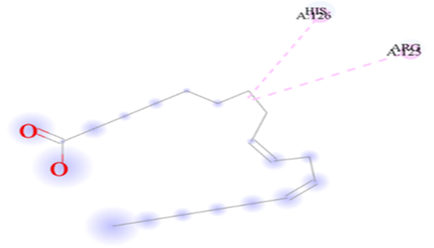
Figure 7.4: The binding of 9,12-Octadecadienoic acid (Z,Z)- against the target macro molecule (DPP4).
Figure 7.5: The binding of n-Hexadecanoic acid against the target macro molecule (DPP4).
Discussion
This study presents integrated evidence on the nutritional value, phytochemical richness, antioxidant potential, antimicrobial activity, and molecular interactions of H. crepitans leaf extract.
The low moisture content suggests improved shelf stability and reduced susceptibility to microbial spoilage [16]. High ash levels reflect significant mineral content, while crude protein and fat values indicate that the leaves can contribute both energy and essential amino acids to the diet [17,18]. The substantial fiber content supports gastrointestinal health by enhancing intestinal motility and reducing metabolic risks [19].
Moderate levels of tannins, oxalates, and phytates were detected. These antinutrients fall within acceptable dietary ranges and are unlikely to impair nutrient absorption under normal conditions, especially when traditional processing methods are applied [20,21].
Mineral analysis revealed appreciable amounts of potassium, calcium, magnesium, and iron, essential for electrolyte balance, skeletal integrity, and oxygen transport [22]. Trace metals were within nutritive ranges, and lead levels were low, indicating minimal toxic risk, although continued monitoring is advisable [4,22].
Phytochemical screening confirmed the presence of saponins, phenolics, alkaloids, anthraquinones, and terpenoids, all associated with antioxidant, antimicrobial, and anti-inflammatory activities [16,19]. Quantitatively, high flavonoid and phenolic contents explain the strong radical scavenging and reducing power observed in DPPH and FRAP assays [8-10].
Antimicrobial activity increased markedly after nanoparticle synthesis, particularly with copper and silver nanoparticles, consistent with reports that nano-formulations enhance membrane interactions and microbial inhibition [23].
Molecular docking supported the experimental findings, with compounds such as gallic acid and estrone showing strong binding affinities to protein targets linked with oxidative stress and metabolic regulation. These interactions provide mechanistic insight into the observed antioxidant and potential antidiabetic effects.
Conclusion
Hura crepitans leaf extract is a rich source of nutrients and structurally diverse bioactive compounds with strong antioxidant and antimicrobial activities. Nanoparticle synthesis significantly enhanced its bioefficacy, while molecular docking confirmed favorable interactions with relevant biological targets. Together, these findings validate traditional uses of H. crepitans and highlight its promise as a source of lead compounds for drug discovery, particularly in oxidative stress and metabolic disorder pathways.
References
- García-González, R., Pérez, E., Martínez, M. (2011). Morphological and ecological characteristics of Hura crepitans and its traditional use in Central America. Revista de Biología Tropical, 59:837-847.
Publisher | Google Scholor - Airaodion, A. I., Ibrahim, A. H., Ogbuagu, U., Ogbuagu, E. O., Awosanya, O. O., et al. (2019). Evaluation of phytochemical content and antioxidant potential of Ocimum gratissimum and Telfairia occidentalis leaves. Asian Journal of Research in Medical and Pharmaceutical Sciences, 7(1):1-11.
Publisher | Google Scholor - Uche, C. L., Ugwu, N. I., Ogbenna, A. A., Okite, U. P., Chikezie, K., et al. (2024). Abnormal Haematological Profile caused by Potassium Bromate in Wistar Rats is corrected by Parkia biglobosa seed. Nigerian Journal of Physiological Sciences: Official Publication of The Physiological Society of Nigeria, 39(1):119-124.
Publisher | Google Scholor - Stevenson, P. C., Green, P. W., Veitch, N. C., Farrell, I. W., Kusolwa, P., et al. (2016). Nor-hopanes from Zanha africana root bark with toxicity to bruchid beetles. Phytochemistry, 123:25-32.
Publisher | Google Scholor - Sofowora, A. (1993). Medicinal plants and medicine in Africa. John Willey Spectrum, Ibadan Nigeria, 281-285.
Publisher | Google Scholor - Trease, G. E., Evans, W. C. (1989). Pharmacognosy. 13th. ELBS/Bailliere Tindall, London, 345-346.
Publisher | Google Scholor - Harborne, A. J. (1998). Phytochemical methods a guide to modern techniques of plant analysis. Springer Science & Business Media.
Publisher | Google Scholor - Gyamfi, M. A., Yonamine, M., Aniya, Y. (1999). Free-radical scavenging action of medicinal herbs from Ghana: Thonningia sanguinea on experimentally-induced liver injuries. General Pharmacology: The Vascular System, 32(6):661-667.
Publisher | Google Scholor - Pulido, R., Bravo, L., Saura-Calixto, F. (2000). Antioxidant activity of dietary polyphenols as determined by a modified ferric reducing/antioxidant power assay. Journal of Agricultural and Food Chemistry, 48(8):3396-3402.
Publisher | Google Scholor - Singleton, V. L., Orthofer, R., Lamuela-Raventós, R. M. (1999). [14] Analysis of total phenols and other oxidation substrates and antioxidants by means of folin-ciocalteu reagent. In Methods in enzymology, Academic Press, 299:152-178.
Publisher | Google Scholor - Bao, J. Y. (2005). Total flavonoids content determination by colorimetric method. Journal of Chinese Medicinal Materials, 28:222-224.
Publisher | Google Scholor - Iqbal, M., Arina, N.B. (2021). Antimicrobial activity of metal nanoparticles. Journal of Nanotechnology, 8859439.
Publisher | Google Scholor - Felpin, F. X., Lebreton, J. (2004). History, chemistry and biology of natural products. European Journal of Organic Chemistry, 18:3693-3712.
Publisher | Google Scholor - Fatoki, T. H., Lawal, O. A., Akinmoladun, A. C. (2018). Phytochemical constituents and bioactivity of medicinal plants. Journal of Medicinal Plants Research, 12(3):45-56.
Publisher | Google Scholor - Badkas, A., De Landtsheer, S., Sauter, T. (2022). Construction and contextualization approaches for protein–protein interaction networks. Computational and Structural Biotechnology Journal, 20:3280-3290.
Publisher | Google Scholor - Esonu, C. E., Iheme, C. I., Njoku, O. C., Agwu, L. O., Dike, C. S., et al. (2024). Assessment of proximate and biomass composition of Cori fiber for potential industrial application. Journal of Nutrition and Food Processing, 7(9):1-6.
Publisher | Google Scholor - Onabanjo, K. S., Airaodion, A. I. (2022). Fortification of “zobo” (Hibiscus sabdariffa) drink with pineapple and watermelon enhances its nutritional qualities and phytochemical compositions. Acta Scientific Nutritional Health, 6(7):3-12.
Publisher | Google Scholor - Esonu, C. E., Iheme, C. I., Njoku, O. C., Agwu, L. O., Edom, C. V., et al. (2024). Investigation of proximate composition and bioactive components in banana (Musa acuminata) peels using advanced analytical techniques. Journal of Nutrition and Food Processing, 7(10):1-9.
Publisher | Google Scholor - Airaodion, A. I., Ayita, E. B., Oluba, S. O., Emaleku, S. A., Osunmuyiwa, O. J., et al. (2021). Chemical composition and nutraceutical potential of velvet tamarind (Dialium guineense wild) fruit pulp. Asian Journal of Biochemistry, Genetics and Molecular Biology, 8(1):1-8.
Publisher | Google Scholor - Ogbuagu, E. O., Ogbuagu, U., Unekwe, P. C., Nweke, I. N., Airaodion, A. I. (2020). Qualitative Determination of the Phytochemical Composition of Ethanolic Extract of Xylopia aethiopica Fruit. Asian Journal of Medical Principles and Clinical Practice, 3(2):159-166.
Publisher | Google Scholor - Airaodion, A. I., Olatoyinbo, P. O., Ogbuagu, U., Ogbuagu, E. O., Akinmolayan, J. D., et al. (2019). Comparative assessment of phytochemical content and antioxidant potential of Azadirachta indica and Parquetina nigrescens leaves. Asian Plant Research Journal, 2(3):1-14.
Publisher | Google Scholor - Airaodion, A. I., Onabanjo, K. S. (2022). Effect of fortifying Zobo (Hibiscus sabdariffa) drink with pineapple and watermelon on the mineral compositions and microbial quality. Merit Research Journal of Food Science and Technology, 7(1):001-007.
Publisher | Google Scholor - Airaodion, A. I., Ekenjoku, J. A., Akaninyene, I. U., Megwas, A. U. (2020). Antibacterial Potential of Ethanolic and Aqueous Extracts of Carica papaya Leaves. Asian Journal of Biochemistry, Genetics and Molecular Biology, 3(3):33-38.
Publisher | Google Scholor