Research Article
Expression Frequency of Esp, Asa373, GelE, CylA and EfaA Genes in Enterococcus Faecalis Clinical Isolates from Biofilm and Planktonic State
- Daniel Duran-Quirós 1
- Eduardo Navas-Barrantes 1
- Diego Ramírez-Forero 1
- Catalina Méndez de la Espriella 1
- Adriana Rodríguez-Ciodaro 1
- Nidia E. de León-Frías 2,3
- Hugo Díez-Ortega 3*
1Faculty of Dentistry, Dental Research Center, Pontifical Javeriana University, Bogotá, Colombia.
2Faculty of Health Sciences, School of Dentistry, Pedro Henríquez Ureña National University, National District, Dominican Republic.
3Infectious Diseases Group, Department of Microbiology, Faculty of Sciences, Pontifical Javeriana University, Bogotá, Colombia.
*Corresponding Author: Hugo Díez-Ortega, Infectious Diseases Group, Department of Microbiology, Faculty of Sciences, Pontifical Javeriana University, Bogotá, Colombia.
Citation: Duran-Quirós D, Navas-Barrantes E, Ramírez-Forero D, de la Espriella CM, Díez-Ortega H, et al. (2025). Expression Frequency of Esp, Asa373, GelE, CylA and EfaA Genes in Enterococcus Faecalis Clinical Isolates from Biofilm and Planktonic State, Dentistry and Oral Health Care, BioRes Scientia Publishers. 4(2):1-7. DOI: 10.59657/2993-0863.brs.25.047
Copyright: © 2025 Hugo Díez-Ortega, this is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Received: June 09, 2025 | Accepted: June 24, 2025 | Published: June 27, 2025
Abstract
The detection and characterisation of virulence genes of Enterococcus faecalis in its sessile (biofilm) and planktonic (free) states is fundamental in endodontics, as it allows a deeper understanding of the mechanisms used by this bacterium to resist treatment, evade the action of antimicrobial agents and the immune system, and persist in root canal infections. Among the main virulence factors associated with the processes of adherence, colonisation, tissue invasion, and biofilm formation are virulence genes, which can modulate their gene expression depending on the environment. The aim of this study was to evaluate the expression of virulence genes Esp, Asa373, GelE, CylA, and EfaA in Enterococcus faecalis isolates of endodontic origin using an “Ex Vivo” model that simulates dental habitat conditions in both sessile and planktonic states. For this purpose, RNA from 63 E. faecalis isolates were RT-PCR amplified for each of the genes. The results showed a higher proportion of positive isolates and high expression of virulence genes in oral mucosa and gutta-percha, with a higher number of cases for Esp, Asa373, EfaA and GelE genes in the sessile state, and CylA in the planktonic state. These data confirm the ability of the bacteria to adhere, form biofilms, invade tissues and resist treatment, raising the need to improve the design of more effective strategies for canal disinfection, the choice of irrigants and intracanal drugs, and the prevention of recurrent endodontic failures.
Keywords: biofilm; enterococcus faecalis; virulence factors; planktonic cells; endodontic infection
Introduction
Enterococcus is a bacterial genus of medical and dental interest that is associated with various community-acquired and hospital-acquired infections, such as urinary tract infections, endocarditis, bacteraemia and sepsis, skin and soft tissue, intra-abdominal, pelvic and biliary tract infections, among others, Enterococcus faecalis is the most representative species [1,2], and in dentistry it is the most frequent bacterium associated with endodontic treatment failures due to its ability to survive in adverse and nutrient-poor environments, with high salt concentrations, high pH and even at temperatures above 45 °C [3]. E. faecalis strains proliferate in both planktonic and forming biofilms, generating a protective barrier that favours their growth and survival in hostile environments [4].
E. faecalis is frequently isolated from the oral cavity, and when the host presents gingivitis, periodontitis, caries and/or pulp necrosis, E. faecalis isolation is common than in healthy patients, and it persists in secondary lesions due to treatment with deficient irrigation, non-localised canals, contaminated instruments, microleakage due to poor obturation and complex anatomies (lateral canals, apical deltas, isthmuses) as well as poor coronal sealing [3,5]. Pathologies at the level of periapical tissues occur under a triad that combines the immunological conditions of the host, the size of the inoculum, the presence of virulence factors and the formation of biofilm in the anatomical site where it is located [3,4,6-8].
Several studies have identified the main virulence factors, the most studied being cell adhesion proteins such as extracellular surface protein (Esp), collagen-binding adhesin (Ace); colonisation facilitator proteins such as aggregation substance (Asa) and its variant Asa373; tissue invasion factors such as cytolysin (CylA), gelatinase (GelE), endocarditis-specific antigen A (EFA), hyaluronidase (Hy); immune evasion factors such as enterococcal polysaccharide capsule (Cps), superoxide ions (O2-), and complementary factors such as lipoproteins, proteolytic enzymes and wall carbohydrates among others [6-15]. Several authors associate the expression of these genes with the different stages of biofilm formation; however, literature differs about the association of the genes with each stage. Despite this, the authors agree that the functionality of the biofilm depends on the strain's ability to adhere to abiotic surfaces, favouring the strain evasion of the host immune system and antimicrobial resistance [12,15,16].
Most studies have shown that E. faecalis isolates derived from different samples of clinical and dental origin have different virulence factor patterns, but that these are based on the presence of the gene when detecting DNA, are evaluated in planktonic stages from pure “In Vitro” cultures, which is why the results are very heterogeneous and sometimes controversial [9,12,17,18]. In this study, to simulate a bacterial ecosystem present in the root canal, an "Ex Vivo" model reproduced in a chamber on extracted human teeth is used to evaluate gene [removed]RNA) and biofilm formation in both sessile and planktonic stages [8].
Materials and Methods
Microbiological Identification
Sixty-three E. faecalis isolates were obtained from ejectors, K-files, mucosal sweep, gutta-percha and paper cones, then cryopreserved at -80 °C until use. Recovery followed the thawed standardised protocol reported by other authors [19]. Briefly, samples frozen in BHI broth with 30 % glycerol (w/v) and supplemented with oxacillin were placed at 4 °C for 30 minutes, followed by another 30 minutes at room temperature. Once thawed, isolates were by simple vortexing homogenised; then 100 uL of each bacterial suspension independently in Enterococci Broth (Becton Dickinson Microbiology Systems, Cockeysville, MD, USA) inoculated and incubated for 24 - 48 hours at 37 °C in Enterococci Chromocult® Enterococci-Agar (Merck, Darmstadt, Germany). Specific species identification by using the MicroScan Pos ID PC34 panel (Dade Behring Inc., West Sacramento, CA, USA) for Gram-positive bacteria followed by reading and interpretation using the autoSCAN-4 system (American MicroScan, Inc., Mahwah, NJ, USA). Each identified isolate was used simultaneously for sessile and plantotechnical culture. Quality control assays were in duplicate, and E. faecalis ATCC 29212 and ATCC 35667 served as controls.
Planktonic Stage
The planktonic culture was by the methodology proposed by Covo et al. (2016). Isolated colonies inoculated in liquid medium Enterococcus Broth (Becton Dickinson Microbiology Systems, Cockeysville, MD, USA) and cultured at 37 ºC for 24 - 48 hours under continuous agitation until a bacterial concentration of 1.5×108 CFU/mL was obtained by spectrophotometer (Thermo Scientific) at OD620 nm [20].
Sessile Stage - "Ex Vivo" Dental Chamber Set-Up
The isolate's capacity to form biofilm in an "Ex Vivo" model was assayed by following a methodology previously established by the research group [8]. One tooth served for each isolate under study. For the preparation of the incubation chamber (in the dental specimen), the crown remotion allows for adjusting all the samples to 16 mm. All the specimens were in 5.25% (w/v) sodium hypochlorite (NaOCl) for 1 minute. The canals were mechanically instrumented with Wave One Goldâ Medium (35.06) (Dentsply-Sirona), within 0.5 mm of the apical foramen, irrigating with 3 mL of 5.25% (w/v) NaOCl as the file advanced through the cervical, middle and apical thirds. Once finished the preparations, final irrigation with 1 mL of 17% (w/v) EDTA for 1 minute, washed with 3 mL of saline solution (0.85% w/v) and dental ducts dried with sterile paper tips (Wave Oneâ, Dentsply-Sirona). The teeth sterilization was at 120 °C for 30 minutes.
For biofilm assembly and formation, 50 mL of each isolate and control strains, at a concentration of 3x108 Cells/mL, were individually inoculated onto the teeth and maintained in a moist, continuous culture medium at 37 °C for 1 week under microaerophilic environment.
Evaluation of the Sessile State Biofilm
The biofilm observation was carried out in scanning electron microscopy and in X-ray energy dispersive spectroscopy. The specimens were sectioned into two parts by cryofracture with liquid nitrogen. Part one was subjected to metallographic preparation with gold for observation in a JEOL scanning electron microscope (SEM), model JSM 6490-LV and analysed under an energy-dispersive X-ray spectroscopy (EDS) detector. The second part of the fractured specimen was eluted with RNA extraction buffer and used to determine the expression of virulence genes associated with the biofilm.
RNA Extraction
RNA extraction from the planktonic cells, occurred once the samples reached a concentration of 1.5×108 CFU/mL by using the RNeasy Mini Kit (Qiagen, Santa Clarita, California). The RNA was in 30 - 100 uL of sterile QP water reconstituted, and concentration spectrophotometrically determined at OD ratio 260/280nm (Nanodrop 2000; Thermo Scientific, Wilmington, DE, USA) [21].
For the sessile state, initially, the methodology proposed by Wang et al. (2011) was followed, in which the bacteria were eluted from the double chamber model [22] and then followed the same protocol used for the planktonic culture.
Retro Transcription-Expression of Virulence Genes
For the two stages, sessile and planktonic, reverse transcription followed the protocol of AccesQuickTM RT-PCR System Kit (Promega-Madison, WI, USA) as described by Covo et al., (2016), 45 ºC for 45 minutes [20]. Amplification protocol followed the conditions by Zhu et al., (2010). Each reaction includes 1 μg of RNA, 10 μL of the Acce AccessQuick™ RT-PCR System commercial kit (Promega), and the primers for the 16S rDNA gene (housekeeping) [18]. Specific primers for Asa/Asa373 (aggregation substance product), Esp (surface protein), CylA (cytolysin), EfaA (Antigen A for E. faecalis) and GelE (gelatinase) were used (Table 1). The PCR products were in 1 % (w/v) agarose gels in 1X TBE buffer solved and analysed on ChemiDocTM MP Imaging System (BioRad Laboratories, Inc). The Molecular Size Marker used was a 1Kb Plus ladder (Invitrogen).
Table 1: Primers used to detect virulence genes [18].
Data Analysis
The data descriptive analysis, as well as gene expression from the two stages, were analysed and compared by Kruskal Wallis statistical. The association between biofilm production and virulence factor expression was analysed using Fisher's test. All statistical analyses were developed using SPSS V15 software (SPSS Inc).
Results
E. faecalis Isolation and Identification
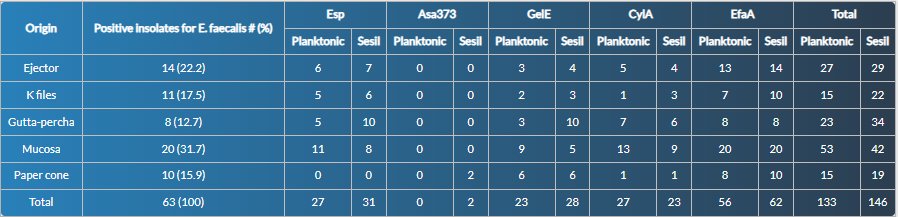
A total of 63 isolates (100 % viable and pure) from the different samples were recovered. The highest number of isolates came from mucosal sweep 31.7 % (20 isolates), ejectors 22.2 % (14 isolates), K-files 17.5 % (11 isolates), paper cones 15.9 % (10 isolates), and gutta-percha 12.7 % (8 isolates) (Table 2).
Table 2: Expression frequency of virulence genes Esp, Asa373, GelE, CylA and EfaA in planktonic and sessile states in Enterococcus faecalis isolates of endodontic origin.
Gene Expression by Stage
After the 63 isolates inoculation in both the dental chamber (sessile) and liquid culture (planktonic), the behaviour observed in the expression of virulence genes was as follows:
All E. faecalis isolates expressed at least one gene; the most expressed gene in both stages (planktonic or sessile) was EfaA (56 and 62 cases). In contrast, Asa373 (0 and 2 cases) was the gene less expressed. The sessile stage showed a higher frequency of expressed genes (146 cases) compared to the planktonic stage (133). A similar situation occurred individually for the genes Esp, Asa373, GelE, and EfaA, with a slightly higher frequency of expressed genes in the sessile stage (Table 2).
Expression of Virulence Genes by Origin
Although no statistically significant differences (p lessthan 0.005) occurred concerning the growth stage or origin of the sample, biologically relevant trends were observed, with a higher frequency of expressed genes in isolates from the oral mucosa, with 53 cases in the planktonic and 42 in the sessile stage (Table 2) and lower frequency of expressed genes in bacterial isolates from the paper cone, with 15 cases in the planktonic and 19 cases in the sessile stage. Similarly, in the mucosa, there was a slightly higher frequency of expressed genes (Esp, GelE, Asa373, and EfaA) in the sessile stage. In contrast, the CylA gene had a higher expressed frequency in the planktonic stage, as well as in mucosal and gutta-percha samples.
Evaluation of Biofilm and Genes in the Sessile State
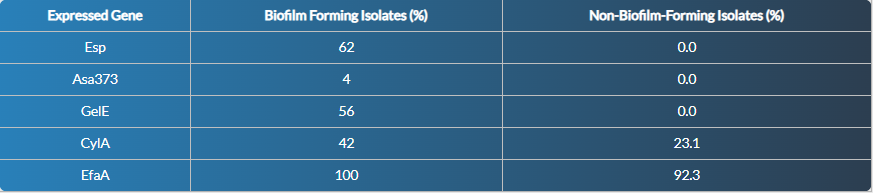
Once analysed the genes expressed in the 63 isolates of the sessile or dental chamber model, we found that Esp and GelE genes were associated with the formation of mature biofilm in E. faecalis, while the CylA gene was not specific for biofilm; EfaA was ubiquitous and not discriminant for biofilm; and Asa373 did not show any association. With these data, the p-value of Fisher's exact test (p lessthan 0.001) indicated that the presence of the Esp and GelE gene was significant and highly associated with biofilm formation.
Table 3: Percentage distribution of genes expressed in the sessile state with respect to biofilm formation in the 63 isolates of E. faecalis.
Discussion
The results of this study showed that E. faecalis is an opportunistic pathogen found in different endodontic source samples such as ejectors, K-files, paper tips and gutta-percha extracted from the obturation of endodontic treatment, with a higher proportion of positive isolates observed in samples from oral mucosa and gutta-percha, suggesting that these environments could act as favourable reservoirs for E. faecalis, allowing its persistence and eventual reinfection of the root canal [22,23].
In its sessile form, E. faecalis develops highly resistant biofilms located in areas inaccessible to conventional mechanical instruments and irrigants. This growth mode confers selective advantages, like increased antimicrobial resistance, immune evasion, and metabolic stability. In the present study, the presence of biofilm and the expression of virulence genes in an "Ex Vivo" model simulating the tooth conditions, was by electron microscopy and RT-PCR evidenced [24-28].
In the comparative analysis of phenotypic states, the Esp, Asa373, EfaA, and GelE genes showed a high expressed frequency in the sessile state, which is consistent with their role (facilitators of cell adhesion, interacting with the extracellular matrix and biofilm formation) [29-31]. In particular, the Esp (enterococcal surface protein) and GelE (gelatinase) genes were significantly associated with the sessile state, reinforcing their role as determinants of E. faecalis virulence [15]. Esp promotes bacterial aggregation and facilitates surface colonisation; some authors have already shown that disruption of this gene affects the bacterial ability to form biofilms [32], while GelE contributes to the degradation of tissue components, favouring invasion and the formation of the biofilm's extracellular matrix.
Gelatinase encoded by the GelE gene is a hydrophobic metalloproteinase with the ability to cleave insulin, casein, haemoglobin, collagen and fibrin. Some studies have associated these proteolytic properties and the high occurrence of Enterococcus species in intraarticular infections, where it has been found to form biofilms on hydroxyapatite [12]. On the other hand, the CylA gene was highly frequently expressed at the planktonic state, which could be due to its involvement in the production of cytolysin, a toxin with haemolytic activity that plays a more relevant role in interbacterial competition and dissemination than in biofilm formation [33].
Asa373, which differs in protein structure from classical Asa, exhibits some moderately conserved amino acid motifs when its sequence is BLAST compared to some other bacterial adhesins. The highest similarity was with the amino acid sequence of the Ag I/II polypeptides of oral streptococci. The Ag I/II polypeptide family mediate collagen recognition in some oral streptococci and is associated with their ability to invade dentinal tubules. However, there is no direct evidence to support the role of Asa373 in mediating collagen binding [12].
The EfaA gene expressed in almost all isolates identified showed a higher expression frequency in the sessile state, supporting its role as a key adhesion factor in surface colonisation during biofilm formation. This behaviour is consistent with previous studies that identify EfaA as an essential element for the interaction of E. faecalis with tissue components, with materials used in root canal filling, and as is known that strains with a higher capacity for biofilm formation had a higher frequency of this gene, suggesting its involvement in the pathogenicity of E. faecalis in persistent infections [34].
This differential pattern of virulence factors in the sessile and planktonic state, as well as in the different samples of endodontic origin, suggests that E. faecalis adapts its virulence profile depending on the environment, maximising its ability to persist by differentially regulating its genes according to its physiological state. These findings are clinically relevant, as biofilm formation is one of the fundamental obstacles to achieving the effective elimination of E. faecalis during root canal treatment. Biofilms protect bacteria from antimicrobial agents, reduce the efficacy of conventional irrigants and hinder intracanal drug penetration [25]. Therefore, the data obtained support the need to incorporate therapeutic strategies aimed at biofilm disruption, the use of more effective irrigants such as activated sodium hypochlorite or EDTA, and the development of new intracanal formulations that include specific anti-biofilm agents.
Finally, the differential expression of virulence genes depending on the phenotypic status and origin of the isolates reinforces the concept of E. faecalis as a highly adaptable pathogenic bacteria, capable of modulating its gene expression to resist adverse conditions and perpetuate endodontic infections. Consequently, it highlights the need to design more effective therapies that include agents specifically directed against biofilm structures or key virulence factors such as Esp, GelE and EfaA, which could be considered therapeutic targets in future research.
Conclusion
Analysis of the behaviour of E. faecalis isolates according to origin and virulence genes inferred that the oral mucosa had the highest proportion of positive isolates and a high frequency of expressed virulence genes, suggesting that it is a key reservoir for E. faecalis. A similar situation occurred in gutta-percha and ejector isolates, which showed a considerable number of virulence genes, pointing to the "probably" persistent colonisation or contamination in dental materials. In contrast, K-files and paper cones showed a lower virulence gene load, signaling a low risk of pathogenicity in the strains from this origin. Meanwhile, the EfaA gene (apparently a constitutive gene) was the most frequent in all samples, reflecting its key role in virulence.
Declarations
Author Contributions
D.D.-Q.; E.N.-B. and D.R.-F.: Conducted laboratory research and wrote the draft manuscript.
C.M.-E. and A. R.-C.: Conceptualized the research and secured the funding for the project.
N.E.L.-F. and H.D.-O.: Conceptualized the research, secured the project funding, conducted part of laboratory research, proofread and approved the final manuscript.
Ethical Statement
All included patients signed an informed consent. The study got the institutional endorsement from the Ethics Committees of the Faculty of Sciences and the Faculty of Dentistry.
Conflict of Interest
The authors declare there are no conflicts of interest. All authors have seen and approved the submitted manuscript. On behalf of all co-authors, the corresponding author shall bear full responsibility for the submission.
Funding
This research was sponsored by Pontificia Universidad Javeriana, project ID: 00007914 - SIAP 00007699 “Caracterización de Enterococcus faecalis de origen endodóntico: perfiles de susceptibilidad antibiótica, expresión de factores de virulencia y actividad ante fagos lítico”.
Acknowledgments
The authors thank the Dental Research Centre at Pontificia Universidad Javeriana, Bogotá, D.C., for supplying the study samples and Dr. Raúl A. Poutou-Piñales, Ph.D., for the English Edition.
References
- Díaz Pérez M, Rodríguez Martínez C, Zhurbenko R. (2010). Fundamental Aspects of The Genus Enterococcus as a Pathogen of High Importance Today. Rev Cubana Hig Epidemiol, 48:147-161.
Publisher | Google Scholor - Saffari F, Dalfardi MS, Mansouri S, Ahmadrajabi R. (2017). Survey for Correlation Between Biofilm Formation and Virulence Determinants in a Collection of Pathogenic and Fecal Enterococcus faecalis Isolates. Infecti Chemother, 49:176.
Publisher | Google Scholor - Pinheiro ET, Mayer MPA. (2014). Enterococcus Faecalis in Oral Infections. J Interdiscipl Med Dent Sci, 3:1000160.
Publisher | Google Scholor - Di Filippo G, Sidhu SK, Chong BS. (2014). The Role of Biofilms in Endodontic Treatment Failure. ENDO, 8:87-103.
Publisher | Google Scholor - Ran S, Gu S, Wang J, Zhu C, Liang J. (2015). Dentin Tubule Invasion by Enterococcus faecalis Under Stress Conditions Ex Vivo. Eur J Oral Sci, 123:362-368.
Publisher | Google Scholor - Ran SJ, Jiang W, Zhu CL, Liang JP. (2015). Exploration of the Mechanisms of Biofilm Formation by Enterococcus faecalis in Glucose Starvation Environments. Aust Dent J, 60:143-153.
Publisher | Google Scholor - Heidari H, Hasanpour S, Ebrahim-Saraie HS, Motamedifar M. (2017). High Incidence of Virulence Factors Among Clinical Enterococcus faecalis Isolates in Southwestern Iran. Infect Chemother, 49:51-56.
Publisher | Google Scholor - Correa Toral F, Delgado Hernández L, Echavarría González C, Serna Varona F, Rodríguez Ciodaro A, et al. (2017). Ex vivo model for studying polymicrobial biofilm formation in root canals. Univ Scient, 22:31.
Publisher | Google Scholor - Zheng JX, Wu Y, Lin ZW, Pu ZY, Yao WM, et al. (2017). Characteristics of and Virulence Factors Associated with Biofilm Formation in Clinical Enterococcus faecalis Isolates in China. Front Microbiol, 8:2338.
Publisher | Google Scholor - Duggan JM, Sedgley CM. (2007). Biofilm Formation of Oral and Endodontic Enterococcus faecalis. J Endod, 33:815-818.
Publisher | Google Scholor - Afonina I, Lim XN, Tan R, Kline KA. (2018). Planktonic Interference and Biofilm Alliance between Aggregation Substance and Endocarditis- and Biofilm-Associated Pili in Enterococcus faecalis. J Bacteriol, 200:e00361-e00318.
Publisher | Google Scholor - Kayaoglu G, Orstavik D. (2004). Virulence Factors of Enterococcus faecalis: Relationship to Endodontic Disease. Crit Rev Oral Biol Med, 15:308-320.
Publisher | Google Scholor - Kafil HS, Mobarez AM, Moghadam MF, Hashemi ZS, Yousefi M. (2016). Gentamicin Induces EfaA Expression and Biofilm Formation in Enterococcus faecalis. Microb Pathog, 92:30-35.
Publisher | Google Scholor - Kafil HS, Mobarez AM. (2015). Spread of enterococcal surface protein in antibiotic resistant Enterococcus faecium and Enterococcus faecalis isolates from urinary tract infections. Open Microbiol J, 9:14-17.
Publisher | Google Scholor - Mohamed JA, Huang DB. (2007). Biofilm Formation by Enterococci. J Med Microbiol, 56:1581-1588.
Publisher | Google Scholor - Ali L, Goraya MU, Arafat Y, Ajmal M, Chen JL, et al. (2017). Molecular Mechanism of Quorum-Sensing in Enterococcus faecalis: Its Role in Virulence and Therapeutic Approaches. Int J Mol Sci, 18:960.
Publisher | Google Scholor - Preethee T, Kandaswamy D, Hannah R. (2012). Molecular Identification of an Enterococcus faecalis Endocarditis Antigen EfaA in Root Canals of Therapy-Resistant Endodontic Infections. J Conserv Dent, 15:319-322.
Publisher | Google Scholor - Zhu X, Wang Q, Zhang C, Cheung GS, Shen Y. (2010). Prevalence, Phenotype, and Genotype of Enterococcus faecalis Isolated from Saliva and Root Canals in Patients with Persistent Apical Periodontitis. J Endod, 36:1950-1955.
Publisher | Google Scholor - Oskouei DD, Bekmen N, Ellidokuz H, Yılmaz Ö. (2010). Evaluation of Different Cryoprotective Agents in Maintenance of Viability of Helicobacter Pylori in Stock Culture Media. Braz J Microbiol, 41:1038-1046.
Publisher | Google Scholor - Covo Morales E, Díaz Caballero A, Simancas Pallares M. (2016). Expression of the Enterococcus faecalis Esp (enterococcus surface protein) Gene in an In Vitro Model of Extracted Teeth. Avances en Odontoestomatología, 32:195-204.
Publisher | Google Scholor - QUIAGEN, (2022). Manual de Instrucciones de uso del RNeasy® DSP FFPE Kit. p.38.
Publisher | Google Scholor - Li W, Yang H, Gong Y, Wang S, Li Y, et al. (2018). Effects of a Chimeric Lysin against Planktonic and Sessile Enterococcus faecalis Hint at Potential Application in Endodontic Therapy. Viruses, 10:290.
Publisher | Google Scholor - Komiyama EY, Lepesqueur LS, Yassuda CG, Samaranayake LP, Parahitiyawa NB, et al. (2016). Enterococcus Species in the Oral Cavity: Prevalence, Virulence Factors and Antimicrobial Susceptibility. Plos One, 11:e0163001.
Publisher | Google Scholor - Chávez de Paz L. (2004). Gram-Positive Organisms in Endodontic Infections. Endod Top, 9:79-96.
Publisher | Google Scholor - Stuart CH, Schwartz SA, Beeson TJ, Owatz CB. (2006). Enterococcus faecalis: Its Role in Root Canal Treatment Failure and Current Concepts in Retreatment. J Endod, 32:93-98.
Publisher | Google Scholor - Yoo YJ, Perinpanayagam H, Oh S, Kim AR, Han SH, et al. (2019). Endodontic Biofilms: Contemporary and Future Treatment Options. Restor Dent Endod, 44:e7.
Publisher | Google Scholor - Quah SY, Wu S, Lui JN, Sum CP, Tan KS. (2012). N-acetylcysteine Inhibits Growth and Eradicates Biofilm of Enterococcus faecalis. J Endod, 38:81-85.
Publisher | Google Scholor - Neelakantan P, Romero M, Vera J, Daood U, Khan A, et al. (2017). Biofilms in Endodontics-Current Status and Future Directions. Int J Mol Sci, 18:1748.
Publisher | Google Scholor - Barbosa-Ribeiro M, De-Jesus-Soares A, Zaia AA, Ferraz CC, Almeida JF, et al. (2016). Antimicrobial Susceptibility and Characterization of Virulence Genes of Enterococcus faecalis Isolates from Teeth with Failure of the Endodontic Treatment. J Endod, 42:1022-1028.
Publisher | Google Scholor - Zoletti GO, Pereira EM, Schuenck RP, Teixeira LM, Siqueira JF, Jr., et al. (2011). Characterization of Virulence Factors and Clonal Diversity of Enterococcus faecalis Isolates from Treated Dental Root Canals. Res Microbiol, 162:151-158.
Publisher | Google Scholor - Tendolkar PM, Baghdayan AS, Gilmore MS, Shankar N. (2004). Enterococcal Surface Protein, Esp, Enhances Biofilm Formation by Enterococcus faecalis. Infect Immun, 72:6032-6039.
Publisher | Google Scholor - Fisher K, Phillips C. (2009). The Ecology, Epidemiology and Virulence of Enterococcus. Microbiol, 155:1749-1757.
Publisher | Google Scholor - Van Tyne D, Gilmore MS. (2014). Friend Turned Foe: Evolution of Enterococcal Virulence and Antibiotic Resistance. Annu Rev Microbiol, 68:337-356.
Publisher | Google Scholor - Ghazvinian M, Asgharzadeh Marghmalek S, Gholami M, Amir Gholami S, Amiri E, et al. (2024). Antimicrobial Resistance Patterns, Virulence Genes, and Biofilm Formation in Enterococci Strains Collected from Different Sources. BMC Infect Dis, 24:274.
Publisher | Google Scholor