Review Article
Challenges and Applications of Artificial Intelligence in Tackling Infectious Diseases and Combating Antimicrobial Resistance
- Aliyu Evuti Haruna 1,2
- Nma Bida Alhaji 2
- John Yisa Adama 3
- Onakpa Michael Monday 1,3
- Hadiza Lami Muhammed 1
- Hussaini Anthony Makun 1
1Africa Centre of Excellence for Mycotoxins and Food Safety Federal University of Technology, Minna, Niger State, Nigeria.
2Livestock productivity and Resilences Support Project, Minna, Niger State, Nigeria, Ministry of Livestock and Fisheries Minna, Niger State, Nigeria.
3Department of Pharmacology and Toxicology, Faculty of Veterinary Medicine, University of Abuja, Nigeria.
*Corresponding Author: Aliyu Evuti Haruna, Africa Centre of Excellence for Mycotoxins and Food Safety Federal University of Technology, Minna, Niger State, Nigeria.
Citation: Aliyu E. Haruna, Nma B. Alhaji, John Y. Adama, Onakpa M. Monday, Hadiza L. Muhammed, et al. (2025). Challenges and Applications of Artificial Intelligence in Tackling Infectious Diseases and Combating Antimicrobial Resistance. Journal of BioMed Research and Reports, BioRes Scientia Publishers. 7(3):1-14. DOI: 10.59657/2837-4681.brs.25.137
Copyright: © 2025 Aliyu Evuti Haruna, this is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Received: February 03, 2025 | Accepted: February 17, 2025 | Published: February 24, 2025
Abstract
Artificial Intelligence (AI) has revolutionized the approach to controlling infectious diseases, significantly accelerating the processes of diagnosis, pathogen detection, and antibiotic discovery. Traditional diagnostic tests often involve lengthy procedures that can delay timely identification of infections, making it challenging to implement appropriate treatments quickly. In contrast, AI technologies, including machine learning and deep learning, have emerged as powerful tools in the realm of medical diagnostics. These advanced systems enable faster and more accurate pathogen detection, predict the resistance profiles of various pathogens, and support the discovery of new drugs, including antimicrobial peptides and small molecules.
AI-driven methods not only enhance the precision of disease diagnosis but also play a crucial role in improving antibiotic stewardship. By predicting which pathogens are likely to develop resistance to certain antibiotics, AI tools help clinicians make better-informed decisions about treatment regimens, ultimately reducing the overuse of antibiotics and mitigating the rise of antimicrobial resistance (AMR). Furthermore, AI aids in identifying promising new drug candidates by analyzing vast amounts of biological data, leading to the discovery of novel compounds that may be effective in combating resistant infections.
This review delves into the multiple applications of AI in the fields of diagnostics, therapeutic interventions, and drug development. It highlights the impressive strengths of these technologies in transforming the landscape of infectious disease management. However, it also acknowledges the challenges and limitations that remain, such as the need for more comprehensive datasets, overcoming biases in AI models, and integrating AI tools into existing healthcare systems. Continued advancements in AI, coupled with collaborative efforts between researchers, clinicians, and data scientists, are essential for further improving the efficiency and effectiveness of AI-driven solutions in tackling the growing threats of infectious diseases and antimicrobial resistance.
Keywords: artificial intelligence; infectious diseases; antimicrobial resistance; diagnosis, pathogen detection; antibiotic
Introduction
In recent years, artificial intelligence (AI) has significantly accelerated the discovery of new antibiotics, achieving in mere hours what traditionally required years of research and development [1–5]. Despite these advancements, infectious diseases remain a major global health challenge, with millions of deaths occurring annually due to antimicrobial resistance (AMR). The widespread misuse and overuse of antibiotics have led to the emergence of pan-drug-resistant pathogens, contributing to an estimated 4 million deaths in 2019 alone [6,7]. However, the failure of antimicrobial treatments is not solely attributable to antibiotic-resistant genes; rather, it is influenced by multiple factors, including the complex biological mechanisms underlying severe clinical conditions such as sepsis [6,8,9]. Addressing this growing public health crisis requires innovative solutions beyond conventional approaches.
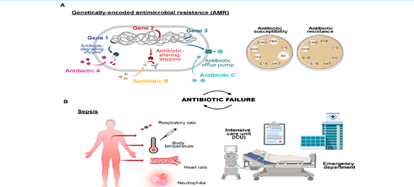
Figure 1: Factors contributing to antibiotic failure: AAntimicrobial resistance driven by genetic exchange or mutation, leading to evolving resistance, and (B) sepsis, a severe response to infection, intensifying the urgency for effective treatment.
The rapid expansion of medical and biological data has fueled the development of advanced AI-driven techniques, enabling computers to process and adapt to increasingly complex information landscapes [10–14]. In the fight against antibiotic failure, AI presents promising opportunities for disease diagnosis and drug discovery, particularly in cases where traditional treatments prove ineffective [15–22) (Figure.1).
Machine learning (ML), a subset of AI, has made remarkable strides across various fields, including computational biology, natural language processing, gaming, and computer vision. Some ML models incorporate deep learning (DL), a sophisticated approach utilizing deep neural networks (DNNs) to identify hidden patterns and make predictions based on intricate datasets, such as biomolecular structures and medical imaging [5,10,23–26]. Cutting-edge AI algorithms have demonstrated the ability to analyze large-scale clinical and experimental data, providing critical insights into disease diagnosis, treatment outcomes, and the discovery of novel therapeutic agents. These advancements underscore AI’s growing role in transforming infectious disease management and enhancing precision medicine [5,10,23–26].
AI-driven technologies have the potential to revolutionize the development of effective antimicrobial compounds, particularly in cases where antibiotic inefficacy threatens patient outcomes. These computer-assisted approaches enable timely and precise interventions, addressing critical clinical conditions that require rapid and accurate decision-making.
This review examines the role of AI-driven methodologies in diagnostics, therapeutic strategies, and drug discovery, providing examples across different levels of application. We discuss the technical aspects and advantages of these approaches, highlighting their ability to enhance efficiency in patient care and treatment development. Furthermore, we address the challenges and limitations associated with these technologies, such as data accessibility and ethical concerns. Emphasizing the importance of standardized datasets and regulatory oversight, we underscore the need for authoritative interventions to ensure data security and ethical compliance in AI-driven healthcare innovations.
Antibiotic failure: an underappreciated global health problem
Antibiotic failure remains a significantly underappreciated global health crisis, primarily driven by the natural selection of antibiotic-resistant phenotypes. A report published in January 2022 identified six major bacterial pathogens Escherichia coli, Staphylococcus aureus, Klebsiella pneumoniae, Streptococcus pneumoniae, Acinetobacter baumannii, and Pseudomonas aeruginosa—as key contributors to millions of deaths worldwide linked to antimicrobial resistance (AMR) [7]. The Centers for Disease Control and Prevention (CDC) has also reported a concerning surge in resistant infections, particularly in hospital settings, positioning AMR among the most pressing threats to public health today [27].
In 2021 alone, AMR was directly responsible for 1.5 million deaths, with an additional 4.71 million deaths associated with resistant infections [6]. A projection published in The Lancet further warns that, if left unaddressed, the burden of AMR could escalate to 1.91 million attributable deaths and 8.22 million associated deaths by 2050. These alarming figures underscore the urgent need for enhanced interventions to mitigate the growing impact of antibiotic resistance on global health.
Figure 2: Using AI to combat antibiotic failure. A Gathering information and data from diagnostic tests, ICU patient clinical conditions, and antimicrobial effectiveness.
B Developing AI-powered models for infection diagnosis, predicting antimicrobial resistance, and tailoring personalized antimicrobial therapies.
The anticipated 10% reduction in antimicrobial resistance (AMR)-related mortality by 2030 is unlikely to be achieved (Fig. 1A). One of the major challenges is the continuous evolution of resistance, combined with the lengthy and complex process of developing new antibiotics. On average, it takes approximately 15 years to bring a new drug to market, encompassing various stages such as discovery, preclinical and clinical trials, regulatory approval by the U.S. Food and Drug Administration (FDA), and post-market safety monitoring [28]. Additionally, the financial burden is substantial, with estimated costs ranging from $161 million to $4.5 billion [29]. Traditional drug discovery relies on high-throughput screenings, which are both time-intensive and costly, as they involve testing numerous compound combinations across different bacterial strains.
The persistent rise of antibiotic-resistant bacteria, coupled with the slow development of effective new antibiotics, is placing significant strain on healthcare systems. As a result, there is growing interest in leveraging advanced artificial intelligence (AI) models and computational techniques to accelerate drug discovery, enabling the identification of novel compounds with desirable properties while expanding the scope of chemical exploration.
Genetically driven AMR contributes to the emergence of resistant strains such as methicillin-resistant Staphylococcus aureus (MRSA), vancomycin-resistant Enterococcus (VRE), carbapenem-resistant Acinetobacter, and extended-spectrum beta-lactamase (ESBL)-producing Enterobacterales. However, genetic resistance is just one factor leading to antibiotic failure. This failure occurs when antimicrobial treatment is no longer effective in eliminating an infection, resulting in persistent or worsening clinical conditions [9]. The mechanisms underlying bacteriostatic activity and drug inactivation are crucial to antibiotic effectiveness; however, additional factors such as pathogen colonization (e.g., biofilm formation) and the body's immune response (e.g., sepsis) also play a significant role in determining treatment outcomes. The type of infection and the patient’s overall clinical condition further influence antibiotic failure [8,9].
Bacterial biofilms, which are structured microbial communities embedded in a polymeric matrix, are responsible for approximately 65% of infections, particularly those involving medical devices such as catheters and implants. Biofilm-associated bacteria exhibit antibiotic resistance levels that are 10 to 1,000 times greater than their free-floating (planktonic) counterparts. This increased resistance is attributed to factors such as restricted antibiotic penetration and altered gene expression [30]. Additionally, biofilms can evade host immune defenses, enabling persistent infections and contributing to chronic disease progression [30]. As a result, early identification of biofilm-forming bacterial species is essential for effective treatment and management [8].
Sepsis, a life-threatening condition triggered by an uncontrolled immune response to infection, often leads to multiple organ dysfunction. In the United States alone, approximately 1.7 million individuals develop sepsis annually, with around 350,000 fatalities occurring during hospitalization [31]. While these figures may not match the projected mortality rates linked to AMR, sepsis-related deaths account for 19.7% of all global fatalities each year [9]. Antibiotics remain the cornerstone of sepsis management; however, their effectiveness is compromised by high mortality rates (ranging from 23% to 35%) and severe clinical complications. Additionally, prolonged or inappropriate antibiotic use exacerbates the problem by fostering the emergence of resistant bacterial strains.
Infections caused by antibiotic-resistant bacteria and the associated adverse side effects often complicate sepsis treatment [32]. The early symptoms of sepsis are typically nonspecific, making it difficult to promptly identify the underlying pathogen and initiate appropriate treatment [9]. Consequently, initial therapeutic approaches tend to rely on empirical broad-spectrum antibiotics, exposing patients to unnecessary antimicrobial agents. This lack of specificity contributes to treatment failure in more than 20% of cases (Fig. 1B) [32].
In this regard, AI-driven methodologies have become valuable in processing vast datasets and uncovering complex patterns that may not be immediately evident to human analysis (Figure. 2A). These technologies support the development of more sophisticated diagnostic tools and enable the creation of personalized treatment strategies tailored to individual patient needs (Figure. 2B) (33).
AI Assistance in Diagnostics
Although artificial intelligence (AI) is still in its early stages for medical diagnosis, the growing availability of data is facilitating its wider application, particularly in disease detection, such as cancer (33). In the case of infectious diseases and sepsis, the complexity of underlying mechanisms and symptom variability poses a significant challenge for early diagnosis. However, machine learning (ML) models show great potential in improving diagnostic accuracy through workflow optimization, task automation, and clinical decision support. Moreover, deep learning (DL) approaches such as convolutional neural networks (CNNs), recurrent neural networks (RNNs), and data mining techniques are playing an increasingly vital role in identifying disease patterns within large datasets (34). These capabilities offer promising advancements in diagnosing, predicting, and classifying diseases across healthcare settings. A summary of various AI models and their potential applications in diagnostics is presented in Table 1.
In intensive care units (ICUs), where patient conditions demand rapid and precise assessments, AI must process diverse data formats, including images, numerical values, and text. Traditional statistical methods, such as linear regression, rely on mathematical equations to establish a "best-fit line," which simplifies complex data relationships. In contrast, DL techniques can analyze intricate, nonlinear relationships more effectively by processing multiple inputs simultaneously, allowing for the development of predictive models tailored to specific medical outcomes.
For instance, Steenkiste et al. utilized a bidirectional long short-term memory (LSTM) model to forecast blood culture results in ICU patients³⁵. LSTMs, a type of RNN, incorporate both current features and previous states as inputs, making them particularly well-suited for analyzing sequential medical data.
AI for Time-Series Data and Microbiology
LSTM models are particularly suited for time-series data due to their ability to retain and selectively forget past information, allowing them to capture long-term dependencies effectively. These models utilize specialized mechanisms to determine which past states to remember or discard, enhancing their ability to track complex relationships over time. In a study by Steenkiste et al., an LSTM-based model was developed to predict blood culture results in ICU patients. The model incorporated nine key clinical parameters temperature, blood thrombocyte count, blood leukocyte count, C-reactive protein concentration, sepsis-related organ failure assessment, heart rate, respiratory rate, international normalized ratio of prothrombin time, and mean systemic arterial pressure—analyzed over time using a high-quality dataset of 2,177 ICU patients. The model demonstrated strong diagnostic performance, achieving an area under the receiver operating characteristic (ROC) curve of 0.99 and an average area under the precision-recall curve (AUC) of 0.82. The ROC curve visually represents a classification model's effectiveness across different threshold settings by plotting sensitivity (true positive rate) against 1-specificity (false positive rate). Meanwhile, the AUC provides a comprehensive metric reflecting the model’s ability to distinguish between positive and negative cases. Notably, the study found that predicting events several hours in advance led to only a minor reduction in accuracy [35].
Beyond clinical diagnostics, AI is increasingly applied in microbiology to analyze data from images, mass spectrometry, and bacterial whole-genome sequencing [16,36]. Smith et al. repurposed a pre-trained convolutional neural network (CNN), initially designed for image classification, to differentiate various bacterial species based on Gram stain morphologies. The model classified Gram-positive cocci in clusters, Gram-positive cocci in chains, and Gram-negative rods from positive blood culture samples. CNNs use multiple layers of filters at different scales to extract detailed features for classification. Their model was trained on approximately 100,000 classified image sections and achieved 95 percentage accuracy in classifying cropped Gram-stain images and 92.5 percentage accuracy for full-slide classifications [37].
Similarly, Zagajewski et al. employed CNNs to distinguish between antibiotic-susceptible and resistant bacterial phenotypes at the single-cell level. This technique enables rapid phenotyping of individual cells exhibiting varied physiological responses to antibiotics [38]. Meanwhile, AI applications using mass spectrometry often compare unknown bacterial proteomic fingerprints expressed as mass-to-charge (m/z) peak values and intensities against a curated database. Traditional software applies predefined rules to assess spectral similarity, whereas machine learning identifies intrinsic data patterns to develop predictive models. This approach has proven especially valuable for antimicrobial resistance prediction, strain typing, and outbreak investigations, eliminating the need for manual programming [16, 39, 40].
Weis et al. compiled mass spectrometry data from clinical bacterial strains, correlating it with resistance profiles to create the DRIAMS dataset. This extensive dataset includes antimicrobial resistance data for over 70 drugs and mass spectrometry information from more than 300,000 clinical isolates across 803 pathogens. Using DRIAMS, three machine learning models—logistic regression, a deep neural network classifier (multilayer perceptron, MLP), and gradient-boosted decision trees (LightGBM)—were trained to predict antimicrobial resistance. Among these, MLP and LightGBM demonstrated the highest classification accuracy⁴¹. This innovative method highlights the potential of AI in enhancing antibiotic stewardship and optimizing treatment strategies efficiently and reliably.
Furthermore, AI and ML techniques are increasingly utilized to analyze bacterial DNA sequences alongside antimicrobial susceptibility data [42]. ML models trained on large libraries of bacterial isolates establish associations between genetic sequences and resistance profiles. Various algorithms, including those analyzing short DNA sequences (k-mers) and assigning predictive weights based on their relevance to antimicrobial resistance, have been developed. Once trained, these models can accurately predict resistance or susceptibility in newly sequenced bacterial isolates, significantly improving infection management and antimicrobial resistance monitoring. Recent studies have shown promising results regarding machine learning (ML) models' ability to predict both the phenotypic profile and the minimum inhibitory concentration (MIC) of antimicrobial agents. For instance, Nguyen et al. utilized an extreme gradient boosting (XGBoost)-based ML model to accurately predict antimicrobial resistance and MIC in nontyphoidal Salmonella [43]. Furthermore, integrating genomic and transcriptomic data can enhance the ability of ML classifiers to predict antimicrobial resistance phenotypes, particularly in Gram-negative ESKAPE pathogens [42].
Alongside high accuracy, the interpretability of these models is essential to foster trust and acceptance among patients and clinicians. There are two primary approaches to explainability: intrinsically interpretable models, also known as "white-box" models, and post-hoc explanations for "black-box" models. In clinical environments, white-box models are often favored because they are easily understood by all stakeholders (44, 45). However, for more complex data, post-hoc methods may be necessary. For example, Drouin et al. developed a sparse, rule-based model, the Set Covering Machine (SCM), to predict antimicrobial resistance using genome k-mers [46]. Meanwhile, Martínez-Agüero et al. combined recurrent neural networks (RNNs) with post-hoc Shapley additive explanations (SHAP) to predict antimicrobial resistance early using ICU electronic health records [47]. SHAP is a model-agnostic tool that ranks features based on their relevance to individual predictions and can be applied across various models (48). Despite its usefulness, post-hoc explainability methods still have limitations, as they only provide imperfect representations of the underlying models. The trade-offs in using these methods must be carefully evaluated for each case [49].
Looking ahead, the number of AI tools, the precision and reliability of AI software analyses, and the integration of AI into clinical microbiology laboratory protocols are expected to grow. Advances in model interpretability will also play a critical role in this progress.
AI in Infectious Disease Treatment
The demand for quick and accessible methods to predict bacterial antimicrobial susceptibility for informed treatment decisions is crucial in clinical environments. AI can significantly contribute to enhancing personalized care by analyzing data such as genetic information, lifestyle factors, and biomarkers, predicting outcomes with targeted interventions, and optimizing clinical treatment strategies.
For example, Kanjilal et al. developed a machine learning (ML) model using electronic health records from 10,053 patients with uncomplicated urinary tract infections (UTIs) in Boston, recorded between 2007 and 2013. This model suggested the most appropriate antibiotics and, when tested retrospectively on 3,629 patients from 2014 to 2016, it outperformed clinicians, reducing the use of second-line antibiotics by 67percentage.
Another notable case was by Yelin et al., who created an ML algorithm to personalize antibiotic prescriptions based on patient demographics, history of urine cultures, and drug purchase records. Using longitudinal data over a decade, they analyzed 711,099 UTI cases involving 315,047 patients. The algorithm successfully predicted antimicrobial resistance tailored to specific drugs, leading to a decrease in mismatched treatments from over 8 percentage to below 6percentage.
In healthcare settings, AI is also utilized in the form of chatbots, which provide patients with timely information about symptoms, prevention, and treatment options through human-like interactions. These chatbots typically consist of four main components: a text comprehension module, a dialog manager, a text generation component, and a database layer for training and operational use. The text comprehension module interprets user inputs using methods like pattern matching (e.g., keyword matching and XML variants like AIML), machine learning techniques (e.g., decision trees and random forests), and natural language processing (NLP) techniques such as named entity recognition. Some chatbots also rely on rule-based systems or fixed input methods, while others use hybrid approaches that combine multiple techniques. For instance, web services like Google's Dialog flow integrate machine learning with rule-based methods. The dialog management module is responsible for processing user inputs and managing the conversation by linking inputs to appropriate responses. There are two main types of data management: static management, which uses pattern matching or rule-based systems, and dynamic management, which leverages machine learning or web services for context switching and identifying intent.
Most chatbots utilize one or more of the following data repositories: a medical knowledge repository (sourced from local or online platforms like Wikipedia), user data storage for response personalization, and conversation scripts, which are often used in pattern matching. By incorporating chatbots into infectious disease management, healthcare providers can improve patient education, enhance access to information, and streamline communication, ultimately leading to better health outcomes. During the COVID-19 pandemic, organizations like the Centers for Disease Control and Prevention (CDC) and the World Health Organization (WHO) adopted chatbots to distribute information, promote safe practices, and offer emotional support. For example, the CDC developed "Clara," a chatbot known as the "coronavirus self-checker," to help individuals determine the appropriate actions if they experience potential COVID-19 symptoms. This chatbot was developed in collaboration with the CDC Foundation and Microsoft's Azure Healthcare Bot service.
Ethics, data privacy, and algorithmic bias
While AI's potential in healthcare is substantial, it also brings significant ethical concerns, including (1) obtaining informed consent for data usage, (2) ensuring safety and transparency, (3) protecting data privacy, and (4) addressing algorithmic fairness and biases. As AI becomes a key component of personalized medicine, safeguarding patient data and ensuring fairness are crucial. The vast amounts of sensitive health information require robust data privacy measures, such as encryption and restricted access, to prevent unauthorized usage. Regulatory bodies like the FDA in the U.S. and the Medicines and Healthcare Products Regulatory Agency (MHRA) in the UK play a vital role in overseeing AI applications in healthcare, ensuring compliance with privacy, safety, and quality standards through audits and inspections. The General Data Protection Regulation (GDPR) enforces strict data privacy requirements, while the European Data Act, effective in January 2024, governs equitable access and usage of data, including health-related data. The WHO emphasizes principles such as transparency, safety, and equity, guiding countries in creating ethical regulations for AI deployment in healthcare and promoting accountability mechanisms.
Moreover, AI models must be built using diverse, representative datasets to prevent the perpetuation of biases in healthcare. Achieving fairness in healthcare means ensuring equal access to care, resources, and outcomes for all patients, regardless of background or social status. In AI-driven healthcare, fairness can be achieved by designing algorithms that are unbiased and capable of providing accurate diagnoses and treatments for all patient groups. Biases may arise from the training data, the design of algorithms, and the interactions between healthcare providers, patients, and AI systems. To address these issues, it is essential to use diverse and representative data, conduct regular audits, and educate both clinicians and patients on recognizing and navigating potential biases.
AI for drug discovery
In addition to its applications in diagnostics and infectious disease treatment, AI-based methods have transformed our ability to predict biomolecular properties and structures, enabling the creation of new active compounds. Machine learning (ML) models, in particular, have provided a way to overcome some limitations of traditional drug discovery approaches, facilitating the rapid design and application of antimicrobial molecules. The growing prevalence of antimicrobial-resistant species and the challenges involved in developing new drugs make it increasingly important to expedite the design of novel antimicrobials.
Antimicrobial peptides (AMPs) and small molecules are both used to treat bacterial infections, but their identification and development have been hampered by the time-intensive nature of traditional experimental screenings and challenges such as toxicity, poor stability, limited cellular penetration, and high synthesis costs. While small molecules are the predominant class of approved antibiotics due to more established research in their design, natural AMPs are emerging as promising alternatives, with potent antibacterial properties and minimal risk of resistance development. Synthetic AMPs also hold potential as future therapeutics, with AMP-based antibiotics like polymyxin B and bacitracin already in clinical use.
Machine learning techniques, such as quantitative structure-activity relationship (QSAR) models, have been used for decades to predict biological activity. These models learn the relationship between input features (such as molecular or peptide descriptors) and biological activity. ML models have been shown to be effective in predicting the biological activity of both small molecule antibiotics and AMPs, and can also predict secondary properties like solubility and toxicity.
Recently, deep learning models have been developed to predict various properties of small molecules and peptides by learning features automatically, rather than relying on predefined molecular descriptors. These models, including graph neural networks (GNNs), encode molecular structures as graphs of atoms and bonds, and can infer global properties based on the hidden states of nodes in the graph. They may also consider three-dimensional molecular features. Additionally, these deep learning models can work with various forms of input, such as predefined descriptors, images, or even natural language descriptions.
The combination of predictive models with graph explainability methods allows for identifying the substructures responsible for specific traits, providing valuable insights into the behavior of molecules. In the search for antimicrobial drugs, these predictive models can be highly effective in virtual screening large databases, enabling rapid identification of potential candidates. For example, they can screen vast genomic and proteomic data for potential AMPs, even searching through billions of possible peptide sequences. Computational analysis has led to the identification of over 2,600 peptide antibiotics within the human proteome and 323 in human gut metagenomes. This computational approach significantly accelerates the process of drug discovery compared to traditional methods.
Figure 3: Using AI to find new antibiotics. Various structural and property descriptions of small molecules and peptides from available databases are used to train predictive and generative AI models.
These models can be trained on different input representations. AI predictions and designs are then tested in wet lab for antimicrobial activity and resistance onset against different pathogens. This workflow can be iteratively executed, where the experimental results from each iteration can inform the training and testing of the AI models in subsequent iterations. Recent advancements in artificial intelligence (AI) have significantly impacted drug discovery, particularly in the identification of new antibiotic molecules. Machine learning (ML) techniques have enhanced our ability to predict biomolecular properties and structures, accelerating the discovery of new compounds (Figure. 3). Traditional drug discovery processes face various limitations, such as time constraints, toxicity issues, and synthesis challenges, making AI-based methods essential for overcoming these hurdles and speeding up the development of antimicrobial agents. Antimicrobial peptides (AMPs) and small molecules both show potential as therapeutic agents, though their clinical translation is often hampered by challenges like toxicity, poor stability, and high synthesis costs. Despite these barriers, AMPs are being increasingly considered as alternatives to traditional antibiotics due to their diverse structures, strong antibacterial activity, and lower likelihood of resistance development. AI-driven approaches have proven effective in predicting the biological activity of both small molecule antibiotics and AMPs, offering a pathway to identify and optimize novel drug candidates more efficiently. Deep learning (DL) models, particularly those based on automatic feature extraction, have shown promise in predicting molecular properties. These models can handle complex representations, such as SMILES strings, amino acid sequences, or even 3D molecular graphs. Graph neural networks (GNNs), which model molecular atoms and bonds as graphs, are particularly useful for predicting the properties of small molecules and peptides. These models can be further enhanced by incorporating three-dimensional molecular features, enabling a more accurate prediction of drug behavior. Additionally, DL methods can speed up in silico virtual screenings, offering a faster alternative to traditional lab-based screenings.
AI has also been used to mine massive biological databases to identify new antimicrobial peptides. For instance, a recent study analyzed over 63,000 metagenomes and 87,000 microbial genomes, discovering nearly a million new antibiotic molecules, many of which demonstrated activity in preclinical mouse models. This comprehensive exploration represents the largest biological data-driven antibiotic discovery effort to date. Moreover, AI-driven models have been applied to search for antibiotic molecules in extinct species, including Neanderthals and Denisovans, marking the beginning of molecular de-extinction. This innovative approach, coupled with deep learning tools like APEX, has uncovered a wealth of potential antibiotic molecules from extinct organisms. These discoveries include several peptides that exhibit potent anti-infective activity, showing the power of AI in uncovering previously inaccessible biological resources. Generative deep learning models are increasingly used to create new molecules by learning from existing biological data and sampling new compounds. These models can be trained on large, unlabeled datasets of drug-like molecules and bioactive peptides, leading to the generation of novel and diverse molecules. Generative models can also be optimized for specific properties, such as antimicrobial activity and low toxicity, by combining reinforcement learning or other optimization strategies. Some AI-generated molecules have shown promising antimicrobial activity, including against resistant bacteria, demonstrating the potential of AI in discovering next-generation antibiotics. However, despite these advancements, challenges remain in AI-driven drug discovery. High-quality labeled data is often scarce or proprietary, making it difficult to train accurate models. Moreover, current models struggle to generate drug candidates that meet the complex criteria required for clinical use, such as toxicity, solubility, and bioavailability. Multimodal models, which combine different data types, are promising solutions but require large, diverse datasets. Finally, understanding and interpreting AI model outputs is essential to avoid errors or false predictions, particularly as these models grow more complex.
Conclusions and Future Perspectives
The continuous advancement of AI technology presents significant opportunities in disease diagnosis and drug discovery, particularly in addressing the challenges posed by antibiotic resistance. AI-driven models, with their capacity to process vast clinical and experimental datasets, are expected to enhance diagnostic precision, predict treatment outcomes, and facilitate the development of novel antimicrobial agents. This transformative potential is particularly evident in critical care settings, where AI can improve diagnostic accuracy and guide personalized treatment plans, ultimately enhancing patient care. Additionally, machine learning (ML) and deep learning (DL) are accelerating the discovery of new antibiotics, thereby strengthening the preclinical antibiotic pipeline—an essential development given the limitations of traditional drug discovery methods and the increasing prevalence of drug-resistant pathogens [110]. Despite these advancements, several challenges persist, particularly regarding data accessibility, which affects the performance and generalizability of AI models across diverse clinical scenarios. Ethical concerns, including issues of informed consent, data privacy, and algorithmic bias, must also be addressed to ensure fair and accurate outcomes. AI models should be trained on diverse and representative datasets to minimize biases that may arise due to imbalanced or incomplete data sources. Many AI-driven approaches for antimicrobial resistance (AMR) diagnosis and treatment rely on datasets that may not fully capture the diversity of patient populations, potentially leading to unreliable or biased predictions [110, 113]. Human cognitive biases and social influences in data collection further exacerbate these issues, making it crucial to adopt high-quality data management and sharing protocols. The quality of AI predictions is highly dependent on dataset composition, where both large datasets with comprehensive details and smaller, well-annotated datasets can be valuable, depending on the specific research objective [110]. For instance, the study of metalloantibiotics remains hindered due to the absence of a publicly available, standardized database, as they are not well-represented using conventional molecular encoding systems such as SMILES [111, 112]. Bridging this gap will require dedicated funding and collaborative efforts to establish robust and standardized datasets, which will, in turn, enhance AI’s capabilities in combating infectious diseases and antibiotic resistance.
Furthermore, AI applications in AMR research are still evolving, and the "black-box" nature of deep learning models raises concerns regarding their interpretability and trustworthiness [113]. Ensuring broader accessibility to high-quality, diverse health data is crucial for developing AI models that are both accurate and reliable across different populations. Limited access to well-rounded datasets restricts algorithm development and perpetuates biases in underrepresented groups. Thus, facilitating data access for researchers will be essential in refining AI-driven diagnosis and treatment strategies, ultimately improving patient outcomes while mitigating algorithmic bias [113].
Looking ahead, we expect the use of AI models that are trained on extensive, high-quality datasets derived from advanced technologies like high-throughput screening and genomic/expression databases. These models will support the development of comprehensive frameworks for quick disease diagnosis and the identification of new antibiotics with novel mechanisms of action. In antibiotic discovery, this involves moving beyond simple proofs-of-concept for hit generation and addressing additional properties, such as toxicity, while incorporating AI into the later stages of drug development. To solve these complex challenges, multiple AI models will need to collaborate and be integrated across various stages of the process. Furthermore, collaboration with healthcare professionals, developers, and regulatory authorities will be crucial for ensuring responsible AI usage, with continuous efforts to uphold fairness and transparency. Regulatory guidelines will play a vital role in protecting patient data and ensuring the safe application of AI.
References
- Wong, F., et al. (2023). Leveraging artificial intelligence in the fight against infectious diseases. Science, 381:164–170.
Publisher | Google Scholor - Cesaro, A., et al. (2023). Deep learning tools to accelerate antibiotic discovery. Expert Opinion on Drug Discovery.
Publisher | Google Scholor - Porto, W. F., et al. (2018). In silico optimization of a guava antimicrobial peptide enables combinatorial exploration for peptide design. Nature Communications, 9:1490.
Publisher | Google Scholor - Liu, G., et al. (2023). Deep learning-guided discovery of an antibiotic targeting Acinetobacter baumannii. Nature Chemical Biology, 19:1342–1350.
Publisher | Google Scholor - Stokes, J. M., et al. (2020). A deep learning approach to antibiotic discovery. Cell, 180:688-702.e13.
Publisher | Google Scholor - Naghavi, M., et al. (2024). Global burden of bacterial antimicrobial resistance 1990–2021: A systematic analysis with forecasts to 2050. The Lancet, 404:1199–1226.
Publisher | Google Scholor - Murray, C. J. L., et al. (2022). Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. The Lancet, 399:629–655.
Publisher | Google Scholor - de la Fuente-Nunez, C., et al. (2023). Antibiotic failure: Beyond antimicrobial resistance. Drug Resistance Updates, 71:101012.
Publisher | Google Scholor - Haney, E. F., et al. (2022). Addressing antibiotic failure—Beyond genetically encoded antimicrobial resistance. Frontiers in Drug Discovery, 2.
Publisher | Google Scholor - Liu, G.-Y., et al. (2024). Antimicrobial resistance crisis: Could artificial intelligence be the solution? Military Medical Research, 11:7.
Publisher | Google Scholor - Ross, J., et al. (2022). Large-scale chemical language representations capture molecular structure and properties. Nature Machine Intelligence, 4:1256–1264.
Publisher | Google Scholor - Ferruz, N., et al. (2024). Anniversary AI reflections. Nature Machine Intelligence, 6:6–12.
Publisher | Google Scholor - Safi, Z., et al. (2020). Technical aspects of developing chatbots for medical applications: Scoping review. Journal of Medical Internet Research, 22:e19127.
Publisher | Google Scholor - Miner, A. S., et al. (2020). Chatbots in the fight against the COVID-19 pandemic. NPJ Digital Medicine, 3:65.
Publisher | Google Scholor - Orsi, M., et al. (2024). Can large language models predict antimicrobial peptide activity and toxicity? RSC Medicinal Chemistry, 15:2030–2036.
Publisher | Google Scholor - Smith, K. P., et al. (2020). Applications of artificial intelligence in clinical microbiology diagnostics. Testing and Clinical Microbiology Newsletter, 42:61–70.
Publisher | Google Scholor - Su, M.-H., et al. (2017). A chatbot using LSTM-based multi-layer embedding for elderly care. Proceedings of the 2017 International Conference on Orange Technologies (ICOT), 70–74. IEEE.
Publisher | Google Scholor - Abashev, A., et al. (2017). Programming tools for messenger-based chatbot system organization: Implication for outpatient and translational medicines. Bionanoscience, 7:403–407.
Publisher | Google Scholor - Orsi, M., et al. (2024). Leveraging machine learning to forecast the antibacterial potential of ruthenium-based compounds. Angewandte Chemie International Edition, 63:e202317901.
Publisher | Google Scholor - Swanson, K., et al. (2024). Utilizing generative artificial intelligence to design and validate structurally novel and easily synthesizable antibiotics. Nature Machine Intelligence, 6:338–353.
Publisher | Google Scholor - Wong, F., et al. (2023). Identifying a new structural class of antibiotics through explainable deep learning methodologies. Nature.
Publisher | Google Scholor - Wan, F., et al. (2024). Applying deep learning for antibiotic discovery via molecular de-extinction techniques. Nature Biomedical Engineering.
Publisher | Google Scholor - Jumper, J., et al. (2021). Achieving highly accurate protein structure prediction using AlphaFold. Nature, 596:583–589.
Publisher | Google Scholor - Zhou, J., et al. (2015). Forecasting the impact of noncoding genetic variants with a deep learning–based sequence model. Nature Methods, 12:931–934.
Publisher | Google Scholor - Rajkomar, A., et al. (2018). Scalable and precise deep learning applications in electronic health records analysis. NPJ Digital Medicine, 1:18.
Publisher | Google Scholor - Centers for Disease Control and Prevention (CDC). (2022). COVID-19: U.S. impact on antimicrobial resistance, special report 2022.
Publisher | Google Scholor - Hutchings, M. I., et al. (2019). A comprehensive review of antibiotics: Historical context, current challenges, and future directions. Current Opinion in Microbiology, 51:72–80.
Publisher | Google Scholor - Schlander, M., et al. (2021). Evaluating the financial burden of drug research and development: A systematic review and cost analysis. Pharmacoeconomics, 39:1243–1269.
Publisher | Google Scholor - Davies, D. (2003). Investigating biofilm resistance mechanisms to antibacterial agents. Nature Reviews Drug Discovery, 2:114–122.
Publisher | Google Scholor - Centers for Disease Control and Prevention (CDC). (2022). What is sepsis?
Publisher | Google Scholor - Silva, B. N., et al. (2013). Evaluating the effectiveness of antimicrobial de-escalation strategies in adult patients with sepsis, severe sepsis, or septic shock. Cochrane Database of Systematic Reviews, 2018:CD007934.
Publisher | Google Scholor - Alowais, S. A., et al. (2023). Transforming clinical practice through artificial intelligence in healthcare. BMC Medical Education, 23:689.
Publisher | Google Scholor - Rabaan, A. A., et al. (2022). The role of artificial intelligence in addressing antimicrobial resistance challenges. Antibiotics, 11:784.
Publisher | Google Scholor - Van Steenkiste, T., et al. (2019). Enhancing the accuracy of blood culture outcome predictions in intensive care units using long short-term memory neural networks. Artificial Intelligence in Medicine, 97:38–43.
Publisher | Google Scholor - Smith, K. P., et al. (2018). Automating the interpretation of blood culture Gram stains using a deep convolutional neural network. Journal of Clinical Microbiology, 56(12):e01521-e01617.
Publisher | Google Scholor - Zagajewski, A., et al. (2023). Leveraging deep learning and single-cell phenotyping for rapid antimicrobial susceptibility detection in Escherichia coli. Communications Biology, 6:1164.
Publisher | Google Scholor - Mather, C. A., et al. (2016). Rapidly detecting vancomycin-intermediate Staphylococcus aureus using matrix-assisted laser desorption ionization-time of flight mass spectrometry. Journal of Clinical Microbiology, 54(4):883-890.
Publisher | Google Scholor - Wang, H.-Y., et al. (2018). Identifying heterogeneous vancomycin-intermediate Staphylococcus aureus through a machine learning-enhanced mass spectrometry approach. Frontiers in Microbiology, 9:2393.
Publisher | Google Scholor - Weis, C., et al. (2022). Predicting antimicrobial resistance directly from clinical MALDI-TOF mass spectrometry spectra using machine learning. Nature Medicine, 28:164-174.
Publisher | Google Scholor - Ali, T., et al. (2023). Artificial intelligence applications for antimicrobial resistance prediction: Challenges and future prospects for real-world implementation. Antibiotics, 12(4):523.
Publisher | Google Scholor - Nguyen, M., et al. (2019). Machine learning to estimate antimicrobial minimum inhibitory concentrations (MICs) and related genomic characteristics for nontyphoidal Salmonella. Journal of Clinical Microbiology, 57(10):e01260-e01318.
Publisher | Google Scholor - Teng, Q., et al. (2022). Interpretability of deep learning in medical diagnostics: A review. Multimedia Systems, 28:2335-2355.
Publisher | Google Scholor - Hakkoum, H., et al. (2022). Systematic mapping and review of interpretability in the medical field. Applied Soft Computing, 117:108391.
Publisher | Google Scholor - Drouin, A., et al. (2016). Reference-free genome comparisons for predictive computational phenotyping and biomarker identification. BMC Genomics, 17:754.
Publisher | Google Scholor - Martínez-Agüero, S., et al. (2022). Interpretable clinical time-series model with intelligent feature selection for early detection of antimicrobial multidrug resistance. Future Generation Computer Systems, 133:68-83.
Publisher | Google Scholor - Lundberg, S., et al. (2017). A unified methodology for interpreting machine learning model predictions. arXiv preprint, arXiv:1705.07874.
Publisher | Google Scholor - Giacobbe, D. R., et al. (2024). Potential and limitations of explainable machine learning in antimicrobial stewardship. Clinical Therapeutics, 46:474-480.
Publisher | Google Scholor - Anahtar, M. N., et al. (2021). Machine learning in addressing antimicrobial resistance: A framework for translational research. Journal of Clinical Microbiology, 59(4):e01260-e01320.
Publisher | Google Scholor - Kanjilal, S., et al. (2020). A decision algorithm to enhance outpatient antimicrobial stewardship for managing uncomplicated urinary tract infections. Science Translational Medicine, 12(545)Leaay5067.
Publisher | Google Scholor - Yelin, I., et al. (2019). Personal clinical history predicts antibiotic resistance in urinary tract infections. Nature Medicine, 25:1143-1152.
Publisher | Google Scholor - Clark, M., et al. (2024). The role of chatbots in healthcare: Facilitating patient access to medical information. Canadian Agency for Drugs and Technologies in Health.
Publisher | Google Scholor - Vijayan, A., et al. (2018). An intelligent learning assistant platform utilizing cognitive computing to support children with autism spectrum disorder. Proceedings of the 2018 International CET Conference on Control, Communication, and Computing (IC4), IEEE, 361-365.
Publisher | Google Scholor - Morris, R. R., et al. (2018). An artificially empathic conversational agent for mental health applications: System design and user perspectives. Journal of Medical Internet Research, 20(6):e10148.
Publisher | Google Scholor - Centers for Disease Control and Prevention (CDC). (2024, April 23). Public health data, surveillance, and technology modernization efforts.
Publisher | Google Scholor - Naik, N., et al. (2022). Legal and ethical considerations of artificial intelligence in healthcare: Responsibility and accountability. Frontiers in Surgery, 9:862322.
Publisher | Google Scholor - Ghanem, M., et al. (2024). AI-driven personalized medicine: Transformative impact on patient care. The New Era of Precision Medicine, Elsevier, 131-142.
Publisher | Google Scholor - Rehman, A. U., et al. (2024). Transforming drug discovery through artificial intelligence advancements. Fundamental Research.
Publisher | Google Scholor - Kadariya, D., et al. (2019). kBot: A personalized, knowledge-driven chatbot for asthma self-management. Proceedings of the 2019 IEEE International Conference on Smart Computing (SMARTCOMP), 138–143.
Publisher | Google Scholor - Aljameel, S. S., et al. (2017). An Arabic conversational intelligent tutoring system to support children with autism spectrum disorder. Proceedings of the 2017 IEEE International Conference on Computational Intelligence and Virtual Environments for Measurement Systems and Applications (CIVEMSA), 24–29.
Publisher | Google Scholor - Martínez-Miranda, J., et al. (2019). User acceptance of a mobile-based embodied conversational agent designed to prevent and detect suicidal behavior. Journal of Medical Systems, 43:246.
Publisher | Google Scholor - Centers for Disease Control and Prevention. (2024). Public health data, surveillance, and technology modernization update.
Publisher | Google Scholor - Ghanem, M., et al. (2024). AI-driven personalized medicine and its transformative impact on patient care. The New Era of Precision Medicine, 131–142.
Publisher | Google Scholor - Bouderhem, R. (2024). The future of AI in healthcare: Ethical and governance perspectives. Humanities and Social Sciences Communications, 11:416.
Publisher | Google Scholor - Ueda, D., et al. (2024). Evaluating fairness in artificial intelligence applications within healthcare: A comprehensive review and strategic recommendations. Japanese Journal of Radiology, 42:3–15.
Publisher | Google Scholor - Chenthamarakshan, V., et al. (2023). Enhancing drug target inhibitor discovery with deep generative foundation models. Science Advances, 9:eadg7865.
Publisher | Google Scholor - Grisoni, F., et al. (2021). Integrating generative AI with on-chip synthesis for de novo drug design. Science Advances, 7:eabg3338.
Publisher | Google Scholor - Merk, D., et al. (2018). AI-driven design of novel bioactive small molecules. Molecular Informatics, 37:1700153.
Publisher | Google Scholor - Zhavoronkov, A., et al. (2019). Rapid identification of potent DDR1 kinase inhibitors using deep learning. Nature Biotechnology, 37:1038–1040.
Publisher | Google Scholor - Das, P., et al. (2021). Accelerating antimicrobial discovery through deep generative models and molecular dynamics simulations. Nature Biomedical Engineering, 5:613–623.
Publisher | Google Scholor - Cesaro, A., et al. (2023). Advanced drug delivery mechanisms for peptide antibiotics. Advanced Drug Delivery Reviews.
Publisher | Google Scholor - Miethke, M., et al. (2021). Sustainable approaches for discovering and developing new antibiotics. Nature Reviews Chemistry, 5:726–749.
Publisher | Google Scholor - Li, J., et al. (2017). Translating mechanistic insights into the design of membrane-active antimicrobial peptides. Frontiers in Neuroscience, 11:73.
Publisher | Google Scholor - Cesaro, A., et al. (2022). Development of synthetic antibiotics from encrypted sequences in human plasma proteins. ACS Nano, 16:1880–1895.
Publisher | Google Scholor - Shatri, G., et al. (2024). Overview of polymyxins. StatPearls.
Publisher | Google Scholor - Nguyen, R., et al. (2024). Bacitracin topical: A review. StatPearls.
Publisher | Google Scholor - Hansch, C., et al. (1962). Examining the correlation between biological activity of phenoxyacetic acids and their physicochemical properties. Nature, 194:178–180.
Publisher | Google Scholor - Huang, J., et al. (2023). Identifying potent antimicrobial peptides through a machine-learning pipeline that explores the full peptide sequence space. Nature Biomedical Engineering, 7:797–810.
Publisher | Google Scholor - Santos-Júnior, C. D., et al. (2024). AI-driven discovery of antimicrobial peptides within the global microbiome. Cell, 187:3761–3778.e16.
Publisher | Google Scholor - Capecchi, A., et al. (2021). Machine learning-assisted design of non-hemolytic antimicrobial peptides. Chemical Science, 12:9221–9232.
Publisher | Google Scholor - Torkamannia, A., et al. (2022). A review of machine learning methodologies for predicting drug synergy in cancer treatment. Briefings in Bioinformatics.
Publisher | Google Scholor - Mayr, A., et al. (2018). Large-scale evaluation of machine learning techniques for drug target prediction using ChEMBL data. Chemical Science, 9:5441–5451.
Publisher | Google Scholor - Bhadra, P., et al. (2018). AmPEP: An AI-based approach for predicting antimicrobial peptides by analyzing amino acid distribution patterns with random forest. Scientific Reports, 8:1697.
Publisher | Google Scholor - Wu, Z., et al. (2018). MoleculeNet: A benchmark dataset for advancing molecular machine learning research. Chemical Science, 9:513–530.
Publisher | Google Scholor - Krenn, M., et al. (2020). SELFIES: A robust molecular string representation ensuring 100% validity. Machine Learning: Science and Technology, 1:045024.
Publisher | Google Scholor - Schütt, K. T., et al. (2018). SchNet: A deep learning framework for modeling molecules and materials. Journal of Chemical Physics, 148:241722.
Publisher | Google Scholor - Ma, Y., et al. (2022). Identifying antimicrobial peptides from the human gut microbiome using deep learning. Nature Biotechnology, 40:921–931.
Publisher | Google Scholor - Jin, W., et al. (2020). Multi-objective molecule generation using interpretable substructures. Proceedings of the 37th International Conference on Machine Learning (ICML 2020), PartF168147-7.
Publisher | Google Scholor - Torres, M. D. T., et al. (2022). Mining the human proteome for encrypted peptide antibiotics. Nature Biomedical Engineering, 6:1451.
Publisher | Google Scholor - Torres, M. D. T., et al. (2024). Investigating human microbiomes uncovers an unexplored reservoir of peptide antibiotics. Cell, 187:5453–5467.e15.
Publisher | Google Scholor - Maasch, J. R. M. A., et al. (2023). Utilizing machine learning to enable the molecular revival of ancient antimicrobial peptides. Cell Host & Microbe, 31:1260–1274.e6.
Publisher | Google Scholor - Cesaro, A., et al. (2023). Artificial intelligence-driven identification of a novel antibiotic. Nature Chemical Biology, 19:1296–1298.
Publisher | Google Scholor - Sterling, T., et al. (2015). ZINC 15 – An accessible ligand discovery platform for researchers. Journal of Chemical Information and Modeling, 55:2324–2337.
Publisher | Google Scholor - Bateman, A., et al. (2023). UniProt: Advancing the Universal Protein Knowledgebase in 2023. Nucleic Acids Research, 51:D523–D531.
Publisher | Google Scholor - Polykovskiy, D., et al. (2020). MOSES (Molecular Sets): A benchmarking framework for molecular generation models. Frontiers in Pharmacology, 11:565644.
Publisher | Google Scholor - Jin, W., et al. (2021). Junction Tree Variational Autoencoder for Molecular Graph Generation. RSC Drug Discovery Series.
Publisher | Google Scholor - Gómez-Bombarelli, R., et al. (2018). Leveraging data-driven continuous molecular representation for automated chemical design. ACS Central Science, 4:268–276.
Publisher | Google Scholor - Gebauer, N. W. A., et al. (2019). Employing symmetry-adapted generation of 3D point sets for targeted molecular discovery. Advances in Neural Information Processing Systems, arXiv preprint arXiv, 32:1906.00957
Publisher | Google Scholor - Jerret, R., et al. (2024). GP-MoLFormer: A foundational model for molecular generation.
Publisher | Google Scholor - Sheth, A., et al. (2007). Transforming big data into smart data: Utilizing semantics to harness volume, variety, and velocity. IEEE Internet Computing, 3.
Publisher | Google Scholor - Irwin, R., et al. (2022). Chemformer: A pre-trained transformer model for computational chemistry. Machine Learning: Science and Technology, 3:015022.
Publisher | Google Scholor - Xie, Y., et al. (2021). MARS: Markov molecular sampling for multi-objective drug discovery. Proceedings of the 9th International Conference on Learning Representations (ICLR 2021).
Publisher | Google Scholor - Guimaraes, G. L., et al. (2017). Objective-Reinforced Generative Adversarial Networks (ORGAN) for sequence generation models.
Publisher | Google Scholor - Zhou, Z., et al. (2019). Enhancing molecular optimization through deep reinforcement learning. Scientific Reports, 9:10752.
Publisher | Google Scholor - You, J., et al. (2018). Goal-directed molecular graph generation using a graph convolutional policy network. Advances in Neural Information Processing Systems.
Publisher | Google Scholor - Yoshida, M., et al. (2018). Exploring sequence space for antimicrobial peptide discovery using evolutionary algorithms and machine learning. Peptide Chemistry, 4:533–543.
Publisher | Google Scholor - Maragakis, P., et al. (2020). A deep-learning perspective on chemical space designed to streamline drug discovery. Journal of Chemical Information and Modeling, 60:4487–4496.
Publisher | Google Scholor - Jin, W., et al. (2019). Learning multimodal graph-to-graph translation for molecular optimization. Proceedings of the 7th International Conference on Learning Representations (ICLR 2019).
Publisher | Google Scholor - Chen, C. H., et al. (2019). Simulation-assisted rational de novo design of a small pore-forming antimicrobial peptide. Journal of the American Chemical Society, 141:4839–4848.
Publisher | Google Scholor - Zhu, Y., et al. (2024). Understanding the coevolutionary dynamics of L2 β-Lactamases using deep learning. Journal of Chemical Information and Modeling, 64:3706–3717.
Publisher | Google Scholor - de la Fuente-Nunez, C. (2024). The role of datasets in artificial intelligence for infectious diseases. Drug Resistance Updates, 73:101067.
Publisher | Google Scholor - Medina-Franco, J. L., et al. (2022). Integrating informatics with medicinal inorganic chemistry: Developing a database for metallodrugs and metallodrug candidates. Drug Discovery Today, 27:1420–1430.
Publisher | Google Scholor - Frei, A., et al. (2023). Utilizing metals in the fight against antimicrobial resistance. Nature Reviews Chemistry, 7:202–224.
Publisher | Google Scholor - Arora, A., et al. (2023). The significance of standardized health datasets in artificial intelligence-driven applications. Nature Medicine, 29:2929–2938.
Publisher | Google Scholor