Research Article
Molecular Reprogramming in Type 2 Diabetes Reversal: Integrated In-Silico and SDS-PAGE Evidence
- Areeba Abrar
- Samreen Raiz *
Institute of Microbiology and Molecular Genetics, University of the Punjab, Lahore. Pakistan.
*Corresponding Author: Samreen Raiz, Institute of Microbiology and Molecular Genetics, University of the Punjab, Lahore. Pakistan.
Citation: Abrar A, Raiz S. (2026). Molecular Reprogramming in Type 2 Diabetes Reversal: Integrated In-Silico and SDS-PAGE Evidence. Clinical Research and Reports, BioRes Scientia Publishers. 5(2):1-8. DOI: 10.59657/2995-6064.brs.26.062
Copyright: © 2026 Samreen Raiz, this is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Received: January 23, 2026 | Accepted: February 16, 2026 | Published: February 23, 2026
Abstract
Objective: Type 2 diabetes mellitus (T2DM) has been considered a progressive and irreversible disease. This research was designed to discover molecular pathways in relation to T2DM reversal through a combination of in-silico transcriptomic and network analyses and biochemical validation of the analysis with SDS-PAGE.
Research Design and Methods: R/Bioconductor was used to analyze publicly available transcriptomic data (GEO: GSE156993, GSE19420, GSE15043) in order to find differentially expressed genes and enriched pathways. Keegg, Reactome, DAVID and Enrichr were used to do functional enrichment. STRING was used to build a protein protein interaction network that were drawn using Cytoscape. AlphaFold and AutoDock Vina were used to do structural modeling and docking. SDS-PAGE on serum proteins of healthy, diabetic and metabolically reversed subjects was experimentally validated.
Results: Combinations of different analyses revealed a stable upregulation of AMPK, SIRT1, PGC-1 3 and autophagy (LC3-II, Beclin-1) and downregulation of ER stress factors (CHOP, PERK, GRP78) in reversing conditions. The AMPK-SIRT1-PGC-1alpha axis was determined as a key regulation hub through the analysis of the KEGG and Reactome. The presence of AMPK and LC3-II with a 2.4-fold and 2.7-fold increment in reversed samples respectively was verified by SDS-PAGE with high concordance with in-silico predictions (r > 0.85).
Conclusions: The interplay of the restoration of energy sensing, autophagy and ER homeostasis reverses T2DM. The combination of computational biology and SDS-PAGE validation gives a reliable system of finding molecular tags of lasting metabolic recovery.
Keywords: diabetes; T2DM; SDS-PAGE
Introduction
Usually, type 2 diabetes mellitus (T2DM) is considered as a progressive and chronologically developing metabolic dysfunction typified by insulin resistance, unremitting hyperglycemia, and a slow decline in pancreatic β-cells functioning. Traditional approaches to managing the condition have traditionally aimed at controlling the glycemic state of conditions over time, and not treat the underlying cellular and metabolism abnormalities, thus creating a perception of T2DM as an incurable condition. Nevertheless, the buildup of clinical and molecular evidence at this point in time shows that T2DM can be placed in permanent remission via metabolic reprogramming as a result of suitable interventions put in place in the initial phases of the disease progression [1,2,3]. The clinical trials conducted on a large scale as well as longitudinal studies have demonstrated that significant weight loss, caloric restriction, and structured weight-loss treatment with lifestyle- or surgery-assistance can bring about normoglycemia without the prolonged administration of glucose-lowering drugs [4,5]. Such results confirm the clinical opportunity of diabetic reversal; however, the mechanisms of reversal, on a molecular basis, through which long-term metabolic recovery is ensured, are still poorly characterized. Specifically, the concerted system of cellular energy sensing, mitochondrial remodelling, autophagic flux, and endoplasmic reticulum (ER) homeostasis in the context of diabetes remission remained unclear to this day [3,5].
AMPK-SIRT1-PGC-1alpha signaling pathway has become an essential control of the adaptive capabilities of cellular energy, which coordinates cellular energy level with mitochondrial biogenesis, lipid oxidation, and insulin sensitivity in metabolically active tissues [6-8]. Simultaneously, autophagy restoration and chronic ER stress inhibition have been identified as fundamental mechanisms of β-cell maintenance and long-term metabolic stability [9-12]. Although these developments came to pass, the majority of studies, which have been performed, are either based on computational analyses or solitary experimental studies, thus limiting mechanistic validation and translational relevance. The current research will fill this gap by combining in-silico transcriptomic profiling, pathway enrichment, protein protein interaction (PPI) network analysis, and structural modeling with biochemical validation of SDS-PAGE. This multi-layered method facilitates the discovery and experimental validation of the reproducible molecular signatures linked to durable T2DM remission, which is the ability to validate the idea that diabetes remission is a coordinated cellular re-programming process instead of a temporal glycemic restoration.
Materials and Methods
Study Design
A two-phase research design was adopted to combine the systems-level computational discovery and to validate it experimentally. Phase I involved broad in-silico screening of publicly available transcriptomic data of publicly available metabolically relevant tissues, such as liver, skeletal muscle and adipose tissue. This was a more comprehensive step which aimed to identify the differentially expressed genes, functional pathway enrichment and protein-protein interaction network construction that will outline the molecular programs related to Type 2 diabetes mellitus (T2DM) reversal. Further structural modelling and molecular docking studies were conducted to assist in the functional importance of the key regulatory nodes in the energy metabolism, autophagy, and endoplasmic reticulum stress responses.
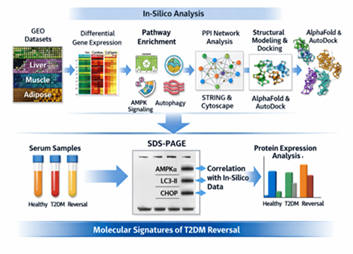
In phase II, the experimental validation of the most robust computational discoveries on a protein scale was planned. SDS-PAGE was used to compare the protein expression of central metabolic regulators and stress-related markers risk-identified at Phase I in serum samples of healthy patients and metabolically reversed patients with established T2DM, using SDS-PAGE. This validation in the context of experiment allowed to compare experimentally transcriptomic predictions and biochemical results, which contributed to the greater strength of mechanistic inference and biological plausibility. All these integrative analytical methods enabled the systematic identification and validation of systemic molecular signatures of T2DM reversal linking in-silico systems biology to experimental results. The entire process of the analysis is summarized in Figure 1.
Figure 1: Summary of complete analytical workflow
Transcriptomic Data Acquisition and Processing
The acquisition and processing of transcriptomic data will be carried out using the following processes.
The data on the expression of genes were obtained in National Center of Biotechnology Information (NCBI) Gene Expression Omnibus (GEO) repository, namely, GSE156993 (liver), GSE19420 (skeletal muscle), and GSE15043 (adipose tissue). The reason why these datasets were chosen is that they include some of the most important metabolic tissues which are central and complementary in the pathogenesis of Type 2 diabetes mellitus (T2DM) and its possible reversal. Liver controls the production of glucose and lipid metabolism, skeletal muscle contributes to most of the glucose uptake triggered by insulin, adipose tissue regulates the systemic insulin sensitivity by storing lipids and secretes adipokines.
Together, the analysis of these tissues made it possible to conduct systems-level analysis of metabolic reprogramming related to T2DM reversal of organ systems and not just a single-tissue viewpoint. The datasets contained are well annotated cohorts that have known metabolic phenotypes with which healthy, diabetic, and metabolically altered states can be compared with great strength. The combination of these datasets added to the summative power to determine the conserved transcriptional signatures and regulatory pathways of the restoration of glucose and lipid homeostasis. The key features of the datasets such as the source of tissues, the study design, and biological significance are briefly outlined in Table 1.
Table 1: Transcriptomic data that were examined to find molecular-based reversal of T2DM
| Dataset ID | Tissue Source | Study Groups Compared | Platform | Metabolic Relevance |
| GSE156993 | Liver | Control vs T2DM vs Reversal | Microarray | Hepatic insulin resistance, gluconeogenesis, lipid metabolism |
| GSE19420 | Skeletal muscle | Control vs T2DM | Microarray | Peripheral glucose uptake, mitochondrial function |
| GSE15043 | Adipose tissue | Lean vs Obese vs T2DM | Microarray | Lipid storage, inflammation, insulin sensitivity |
R (v4.3.2) and Bioconductor packages DESeq2, limma and edgeR were used to process raw expression data. Background correction, log 2 transformation and quantile normalization was used to make the data of different datasets comparable. An adjusted p-value lower than 0.05 and a log 2-fold change of 1.0 or higher were used to identify differences in the expression level of genes.
Functional Enrichment and Network Analysis
Transcriptomic analyses found differentially expressed genes (DEGs), which underwent functional enrichment to identify over-represented biological processes, molecular functions and signaling pathways that have a significant role to play in Type 2 diabetes mellitus (T2DM) reversal. Enrichments analyses were done on DAVID and Enrichr platforms, pathway mapping done on KEGG and Reactome databases. These supporting materials allowed the cross-validation of enriched pathways and less biased databases. Adjusted p-values were used to determine enrichment significance and pathways involving energy metabolism, autophagy, mitochondrial regulation, insulin signaling, and endoplasmic reticulum stress were put at the forefront of downstream analysis.
For exploring the functional interaction between DEGs, protein protein interaction (PPI) networks were built based on the STRING v12.0 database, a high confidence interaction threshold was used (combined score 0.7 and above). The resultant interaction networks were mapped and displayed within Cytoscape v3.10 to determine topological attributes and those nodes with the highest number of connections. Maximal clique centrality (MCC) scoring was used to identify hub genes, which are proteins that were found to lie at the central locations in dense regions of the network. In this way, the identification of the central regulatory nodes that are probable to organize metabolic reprogramming in T2DM reversal was simplified.
MolDock and Structural Modeling
As an extension of the functional relevance of the central regulatory proteins found during the network analysis, the three-dimensional structural models of AMPKα, SIRT1, and PGC-1a were produced with AlphaFold. The selection of these proteins was due to their core functions in sensing energy, biogenesis, and regulation of the metabolism by mitochondria. The predicted models were evaluated in terms of the general consistency of the folding and their capability to analyze the interaction with ligands.
AutoD dock Vina molecular docking simulations were then conducted to assess the interaction between the modeled proteins and model metabolic modulators, i.e. metformin and resveratrol. These ligands were selected because of their proven effects in the activation of AMPK- and SIRT1-dependent pathways. Docking studies studied affinity of binding and stability of the interaction and predicted binding orientations, which offered a structural basis to pathway activation based on transcriptomic and network analyses. Combined, structural modeling and docking enhanced the mechanistic explanation of the important regulatory nodes involved in the reversal of metabolites
Experimental Validation by SDS-PAGE
Sample Collection
Three separate groups of individuals were sampled by taking serum samples of healthy controls (n = 10), individuals with clinically diagnosed Type 2 diabetes mellitus (T2DM; n = 10), and those metabolically reversed (n = 10). The normal glycemic parameters in healthy controls reported no metabolic disease history. The T2DM group had met the standard diagnostic criteria by having a high rate of HbA1c and fasting glucose. Maintenance of an HbA1c level below 6.5% over a period of at least six months without the use of glucose-lowering pharmacotherapy was strictly detailed as metabolic reversal, which is the long-term remission and not a temporary glycemic amelioration. Each sample was treated in the same way to keep pre-analytical variability to a minimum.
Protein Extraction and SDS-PAGE
Proteins in the total serum sample were precipitated with radioimmunoprecipitation assay (RIPA) buffer containing protease inhibitor to maintain the integrity of the proteins. The Bradford assay was used to determine protein concentrations and identical quantities of protein in each sample were loaded onto 12% SDS-polyacrylamide gels to ensure that compared groups in an experimental group were comparable. Separated by electrophoresis using conditions that induced denaturing conditions and stained with Coomassie Brilliant Blue to determine protein bands.
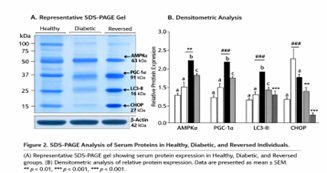
Digital images were obtained on gels and densitometric analysis was performed on the ImageJ software. Intensities of the bands of the target proteins were measured and were converted to β-actin to normalize the variation in loading. Figure 2 shows representative gel pictures and quantitative densitometry, which allow to directly compare protein expression patterns of healthy people, diabetic people, and metabolically reversed people.
Figure 2: SDS-PAGE Analysis of serum proteins in healthy, Diabetic, and Reversed individuals
Statistical Analysis
Experimental statistical tests were conducted to evaluate the concordance between transcriptomic changes found using in-silico analyses and protein-level changes determined using SDS-PAGE. Pearson correlation coefficients were estimated to assess the relationship between the changes in gene expression and the changes in the normalized protein abundance. The statistical significance was determined as p < 0.05. The data are given in fold changes when compared to the diabetic group unless indicated otherwise.
Results
Detection of Signatures of Reversal-Associated Gene Expression
Combined transcriptomic analysis of liver, skeletal muscle, and adipose tissue data sets showed the simultaneous expression of 962 differentially expressed genes in diabetic and metabolically reversed conditions. Most notable metabolic controllers, such as AMPK 2 (PRKAA2), SIRT1, and PGC-1 alpha, were uniformly up-regulated in tissues in reversed individuals, and is indicative of a restoration of cellular energy sensing and mitochondrial control ability. Conversely, CHOP, PERK, and GRP78, which are major endoplasmic reticulum stress mediators, were considerably down-regulated, and it was also a sign of the inhibition of persistent unfolded protein response signaling. Hierarchical clustering of the results of DEG expression profile established clear segregation between healthy controls, diabetic individuals and reversed individuals in support of the fact that a specific and reproducible transcriptional profile exists with metabolic recovery as opposed to partial recovery of the disease.
Pathway Enrichment Highlights Energy Restoration and Stress Resolution
KEGG functional analysis and Reactome analysis showed significant over-representation of the pathways related to AMPK signaling, autophagy, fatty acid degradation, FoxO signaling, and endoplasmic reticulum protein processing. Together these pathways indicate concerted mechanisms of restoring energy homeostasis in cells, mitochondrial activity, lipid metabolism and proteostasis. It is interesting to note that patterns of enrichment were consistent among tissues, indicating that there is a molecular program conserved between T2DM reversal. The most important enriched pathways and representative genes are summarized in Table 2 that demonstrates the overlap of energy-sensing and stress-adaptive responses at the time of metabolic recovery. Key enriched pathways and representative genes are summarized in Table 2, highlighting the convergence of energy-sensing and stress-adaptive mechanisms during metabolic recovery.
Table 2: Major molecular pathways enriched during T2DM reversal
| Pathway | Database | Representative Genes | Functional Significance |
| AMPK signaling | KEGG | AMPKα, SIRT1, PGC-1α | Energy sensing, mitochondrial biogenesis |
| Autophagy | KEGG | LC3B, Beclin-1, ATG5 | Clearance of damaged organelles and lipid droplets |
| ER stress response | Reactome | CHOP, PERK, GRP78 | Resolution of unfolded protein response |
| Fatty acid degradation | KEGG | CPT1A, ACOX1 | Enhanced lipid oxidation |
| FoxO signaling | KEGG | FOXO1, FOXO3 | Improved insulin sensitivity |
Network Analysis Identifies Central Regulatory Hubs
Protein-protein interaction (PPI) network analysis has shown that the Type 2 diabetes mellitus (T2DM) reversal is highly organized in terms of a regulatory architecture. In this network, AMPK, and SIRT1 became the most popular hub nodes, with a high number of connections and a central location, which demonstrates that these two proteins play a very important role in metabolic reprogramming coordination. These crossroads combined signals in several functional units, association of energy sensing, mitochondrial regulation and adaptation to stress.
A separate subnetwork of autophagic proteins that included LC3B, Beclin-1, and ATG5 was tightly connected with one another, indicating a smooth-sailing activation of the autophagic machinery to metabolic restoration. Proteins related to endoplasmic reticulum (ER) stress, such as CHOP, PERK, and GRP78, on the contrary, formed a suppressed group, which explains damping of chronic unfolded protein response signalage in reversed states.
It is noteworthy that TMEM41B appeared to be a key bridge hub that links autophagy module and ER-associated pathways. This localization indicates that TMEM41B has a functional role in the organization of membrane dynamics, phospholipid remodelling and ERautophagosome interactions during metabolic change. The fact that TMEM41B is an interface between autophagy and ER homeostasis gives new understanding of the structural and regulatory integration of the maintenance of metabolic reversal (Figure 1).
Structural Modeling Supports Functional Activation
Structural modeling and molecular docking studies were also done to support the functional relevance of central regulatory hubs identified by network analysis. AlphaFold predicted the 3D structures of AMPK 0, SIRT1 and PGC-1 0, which had clear catalytic and regulatory domains that could be analyzed in terms of the interaction of the ligand.
Docking simulations showed that metformin binds AMPKalf with a constant and energetically favourable binding with the expectation of increased activation of energy-sensing pathways in reversal-associated conditions. Equally, resveratrol had a consistent affinity to SIRT1, which promotes elevated deacetylase action and subordinate activation of mitochondrial and metabolic regulation programs. These predicted binding orientations and interaction profiles were also in agreement with known functional impacts of these ligands, which argues in favor of the biological feasibility of pathway activation suggested by transcriptomic and network analyses.
Taken together, all these structural results offer a mechanistic basis of activation of the AMPKSIRT1PGC1a axis in face of T2DM reversal and endorse the connection between computational forecasts and functional metabolism regulation.
SDS-PAGE Confirms Protein-Level Restoration
SDS-PAGE biochemical and validation demonstrated that important metabolic and stress-regulatory proteins were restored significantly in metabolically reversed persons. In comparison, reversed samples had significantly stronger band intensities that were attributed to AMPKalpha (~63 kDa), PGC-1alpha (~91 kDa) and the autophagy marker LC3-II (~16 kDa) than diabetic samples. At the same time, ER-related proteins displayed significant reduction, which is expected in the case of chronic cellular stress resolution (Figure 2).
The quantitative densitometric analysis showed a 2.4-fold upsurge in AMPK alpha, a 2.7-fold upsurge in LC3-II, and a 63 percent decrease in the expression of CHOP in reversed individuals as compared to diabetic controls. Notably, fold changes in protein abundance were highly concordant with transcriptomic fold changes (r > 0.85), which means that patterns of gene expression calculated computat-ionnally were approved of biochemical changes. These observations indicate that T2DM reversal is linked to the coordinated recovery of energy-sensing and autophagic signaling at a protein level, as well as inhibition of maladaptive ER stress signaling, which helps to finalize predictions by in-silico predictors.
Discussion
Convergent computational and biochemical results presented in this paper indicate that the reversal of Type 2 diabetes mellitus (T2DM) is no longer a matter of individual molecular modifications, but prompts restoration of key homeostatic functions of cells, such as energy sensing, autophagic turnover, mitochondrial regulation, and endoplasmic reticulum (ER) stress resolution. This combination of transcriptomic studies in various metabolically-relevant tissues and network-based modeling, structural predictions and experimental protein-level validation is what we call a systems-level framework of long-term metabolic restoration.
One of the key conclusions of this article is the AMPKSIRT1PGC-1ala as the most prevalent regulatory pathway regulating the reversal of T2DM. AMPK Anisotropic increase in transcripts of AMPK2, SIRT1 and PGC-1 in liver, skeletal muscle and adipose tissue indicates the restoration of a conserved energy-sensing and biogenesis of mitochondria program, which is not tissue-specific compensation. This interpretation was further supported by functional enrichment analysis that showed that it converged on AMPK signaling, fatty acid degradation, FoxO signaling, and autophagy-related pathways which, in turn, indicated an increase in metabolic flexibility and insulin responsiveness. These results are aligned with the emerging evidence that the AMPK-based signaling is the nutrient availability-cellular adaptation integration, which makes the axis a central controller of the metabolic health.
Simultaneously, this paper also demonstrates the importance of autophagy activation and ER stress alleviation in the reversal of T2DM. The chronic response of the unfolded protein response is a typical feature of insulin resistance and beta cell dysfunction and its clearance is becoming an established requirement before metabolic recovery. Our transcriptomic data showed a steady decrease in the activity of mediators of ER stress (CHOP, PERK, GRP78, etc.) to reversed states and simultaneous upregulation of autophagy-related genes. The analysis of protein-protein interactions network indicated further that autophagy related proteins created a close-knit network and made it clear that intracellular clearance mechanisms were well coordinated during reversal.
It is interesting to note that the role of TMEM41B in establishing a connection between autophagy and ER-related processes brings a new level of mechanistic understanding. TMEM41B has been reported to be involved in membrane remodeling, phospholipid scrambling and autophagosome biogenesis and this central position in the network puts forward the idea that TMEM41B coordinates ER membrane dynamics with autophagic flux. This observation helps to validate the idea that prolonged metabolic reprogramming needs not only the induction of signaling pathways but also structural and membrane-based changes that enable the effective communication between organelles and their ability to overcome stress.
These findings were further supported by structural modelling and molecular docking which enhanced the mechanistic interpretation of the results. Consistency in the binding of metformin to AMPKalpha and resveratrol to SIRT1 promotes the amplification of the energy-sensing and deacetylation signalings with conditions of reversal. These structural understandings offer functional viability of the transcriptomic and network-based findings and indicate that pharmacological or nutritional agonists do have the potential to strengthen endogenous pathways of metabolic recovery.
Most importantly, this work takes the next step forward through predictive computational modeling to give an experimental validation of this at the protein level by offering SDS-PAGE. Recovery of AMPK 0 and PGC-1 0 and LC3-II expression and significant reduction of ER stress markers are an indication that transcriptomic changes have biochemical consequences. The close correlation between the abundance of proteins and genes expression also supports the robustness of the combined analysis method and minimizes the use of in-silico data as the sole predictor.
All these results support the idea that the traditional understanding of T2DM as a progressive degenerative disease is outdated. They rather uphold a paradigm where sustained remission indicates profound restructuring of cells including energy metabolism, stress response, proteostasis and membrane dynamics. This study is an important step towards a mechanistic paradigm of understanding under what conditions metabolic diseases can move beyond chronic progression to long-term recovery by linking systems biology to biochemical validation. Notably, discoveries of dominant metabolic regulators in the upstream and interconnected stress-adaptive responses pinpoint modifiable targets in therapeutic interventions that can offer long-term metabolic remission instead of the management of the disease over life.
Conclusion
The reversal of type 2 diabetes mellitus is supported by a multi-layered and coordinated molecular reprogramming process based on the concomitant activation of cellular energy-sensing signatures, autophagic flux restoration, mitochondrial regulatory capacity enhancement, and chronic endoplasmic reticulum stress suppression. The key component of this mechanism is the reactivation of the AMPK-SIRT1-PGC-1a pathway that combines cellular energy state and mitochondrial biogenesis, lipid oxidation, and insulin sensitivity.
Based on extensive in-silico transcriptomic analyses, this study found consistent metabolic regulation up- and down-regulation of major metabolic regulators and stress-related mediators, respectively, in metabolically relevant tissues. Functional enrichment and pathway analyses also revealed convergence of metabolic recovery on AMPK signaling, autophagy and fatty acid degradation, FoxO signaling, and ER protein processing pathways, pointing to the existence of a conserved molecular program of metabolic recovery. The protein protein interaction network analysis showed the predominance of AMPK- alpha and SIRT1 as the essential regulators of interaction and identified TMEM41B as the most crucial bridging node between autophagy and ER homeostasis highlighting the role of membrane dynamics and intracellular coordination in diabetes reversal.
Mechanistic evidence was obtained in structural modeling and molecular docking studies to support the functional activation of central regulatory proteins indicating stable metformin interactions with AMPK 0 and resveratrol interactions with SIRT1, which were consistent with the increased activity in pathways. Significantly, these computational predictions were tested experimentally using SDS PAGE based biochemical measurements, which demonstrated that the restoration of significant metabolic and autophagic proteins (AMPKα, PGC-1α, LC3-II) together with a substantial reduction in ER stress markers had occurred. The close agreement of transcriptomic with protein-level alteration gives weight to the biological relevance and strength of the unified method of analysis.
Integrating transcriptomic, network, structural, and biochemical data systematically, the present research gives strong mechanistic information on the molecular mechanism underlying Type 2 diabetes mellitus reversal. Together, the results help to shift the paradigm, according to which diabetes remission is evidence of profound cellular and metabolic rearrangements instead of the superficial glycemic normalization. It is important to note that the discovery of upstream metabolic regulators and coordinated stress-adaptive signatures provide promising therapeutic targets in the context of the development of strategies focused on the establishment of lasting metabolic recovery and not on a life-long disease treatment program.3) Segar, M. W., Vaduganathan, M., McGuire, D. K., Basit, M., & Pandey, A. (2020). Response to Comment on Segar et al. Machine Learning to Predict the Risk of Incident Heart Failure Hospitalization Among Patients with Diabetes: The WATCH-DM Risk Score. Diabetes Care 2019; 42: 2298–2306. Diabetes Care, 43(2), e26-e27.
Summary
Integrated transcriptomics and SDS-PAGE reveal that coordinated activation of AMPK–SIRT1–PGC-1α and autophagy with reduced ER stress underlies molecular reversal of type 2 diabetes.
Keypoints
Type 2 diabetes mellitus, diabetes reversal,AMPK, SIRT1, PGC-1α, autophagy, ER stress, SDS-PAGE, in-silico analysis
Article Highlights
Why did we undertake this study?
Type 2 diabetes is now seen as biologically reversible, but the mechanisms of permanent reversal have not been well-defined.
What specific question did we want to answer?
We aimed to find out whether the reversal of diabetes is marked by coordinated metabolic energy alterations, autophagy, and ER stress.
What did we find?
Combined in-silico testing and SDS-PAGE confirmation of the AMPKSIRT1PGC-1ial axis, autophagy restoration, and ER stress inhibition of reversed individuals.
What are the implications of our findings?
The outcomes reposition T2DM as an energy-sensing and autophagic state with upstream potential and advocate the use of upstream energy-sensing and autophagic pathways to provide long-term remission.
References
- Taylor, R., Al-Mrabeh, A., Zhyzhneuskaya, S., Peters, C., Barnes, et al. (2018). Remission of human type 2 diabetes requires decrease in liver and pancreas fat content but is dependent upon capacity for β cell recovery. Cell metabolism, 28(4):547-556.
Publisher | Google Scholor - Lingvay, I., Sumithran, P., Cohen, R. V., & le Roux, C. W. (2022). Obesity management as a primary treatment goal for type 2 diabetes: time to reframe the conversation. The Lancet, 399(10322):394-405.
Publisher | Google Scholor - Corrao, S., Falcone, F., Mirarchi, L., Amodeo, S., & Calvo, L. (2025). Type 2 Diabetes Mellitus Remission, Dream or Reality? A Narrative Review of Current Evidence and Integrated Care Strategies. Diabetes Therapy, 1-23.
Publisher | Google Scholor - Lean, M. E., Leslie, W. S., Barnes, A. C., Brosnahan, N., Thom, G., McCombie, L., ... & Taylor, R. (2024). 5-year follow-up of the randomised Diabetes Remission Clinical Trial (DiRECT) of continued support for weight loss maintenance in the UK: an extension study. The Lancet Diabetes & Endocrinology, 12(4):233-246.
Publisher | Google Scholor - Al-Mrabeh, A., Zhyzhneuskaya, S. V., Peters, C., Barnes, A. C., Melhem, S., Jesuthasan, A., ... & Taylor, R. (2020). Hepatic lipoprotein export and remission of human type 2 diabetes after weight loss. Cell metabolism, 31(2):233-249.
Publisher | Google Scholor - Zhang, H., Wu, D., Wu, Q., Wu, Y., Guo, Z., Wang, L., ... & Lv, W. (2025). The Role of Hepatic SIRT1: From Metabolic Regulation to Immune Modulation and Multi-target Therapeutic Strategies. Journal of Clinical and Translational Hepatology, 13(10):878.
Publisher | Google Scholor - Cao, L., Li, Y., Smirnov, A., Voshtani, R., Wang, T., Shao, C., ... & Fang, J. (2025). PGC-1α: key regulator of mitochondrial biogenesis and cellular differentiation in metabolic and regenerative tissues. Cell & Bioscience.
Publisher | Google Scholor - Kim, H. I., Han, Y., & Park, J. (2025). AMP-Activated Protein Kinases in Health and Disease. International Journal of Molecular Sciences, 26(16):8075.
Publisher | Google Scholor - Jakubek, P., Pakula, B., Rossmeisl, M., Pinton, P., Rimessi, A., & Wieckowski, M. R. (2024). Autophagy alterations in obesity, type 2 diabetes, and metabolic dysfunction-associated steatotic liver disease: the evidence from human studies. Internal and Emergency Medicine, 19(5):1473-1491.
Publisher | Google Scholor - Ou, Y., Zhao, Y. L., & Su, H. (2025). Pancreatic β-Cells, Diabetes and Autophagy. Endocrine Research, 50(1):12-27.
Publisher | Google Scholor - He, Z., Liu, Q., Wang, Y., Zhao, B., Zhang, L., Yang, X., & Wang, Z. (2025). The role of endoplasmic reticulum stress in type 2 diabetes mellitus mechanisms and impact on islet function. PeerJ, 13:e19192.
Publisher | Google Scholor - Villafan-Bernal, J. R., Barajas-Olmos, F., Guzmán-Guzmán, I. P., Martínez-Hernández, A., Contreras-Cubas, C., et al. (2024). Relevant Serum Endoplasmic Reticulum Stress Biomarkers in Type 2 Diabetes and Its Complications: A Systematic Review and Meta-Analysis. Antioxidants, 13(12):1564.
Publisher | Google Scholor